Space Ranger6.4, printed on 03/23/2025
The Space Ranger pipeline uses automatic image detection algorithms to calculate the location of the fiducial markers, and find the boundaries of the tissue. 10x Genomics tested the algorithms on a wide variety of H&E images, tissues and samples, and found the detection algorithms to be robust. However, for fluorescence images, or in cases where the fiducial markers are obscured or tissue boundaries are unclear, manual fiducial alignment and tissue outlining will be required.
Loupe Browser 4.0 onwards provides the Visium Manual Alignment wizard for fluorescence images and other cases where more manual intervention is required. The alignment wizard makes it possible to interactively align an image to a slide’s Fiducial Marker locations, perform fine-grained tissue selection with a suite of tools, and export the manual alignment into a Space Ranger run. This tutorial walks through this process step-by-step.
Loupe Browser 5.1 onwards includes the option to load an additional overexposed capture to facilitate fiducial selection for fluorescence images. This overexposed capture can be included as an additional channel in the same image file, or as a separate image file, depending on imaging system capabilities. This capture is excluded from downstream analysis in Space Ranger and Loupe Browser.
Loupe Browser 6.2 onwards multiple aligner windows can be simulatanouesly opened for independent image analysis workflows.
The tutorial below demonstrates the alignment steps for a single-channel brightfield image, and a multi-page fluorescence image for steps that differ.
The Visium Manual Alignment wizard can be found on the Loupe Browser start page, at the bottom of the main panel. Clicking on the link starts the alignment wizard in a new window. Since, Loupe Browser 6.2 introduced an additional image alignment workflow to support the CytAssist Image Alignment, the landing page now offers the two workflow choices. Click on the box corresponding to Fiducial Alignment.
This opens a pop-up window listing the steps required for completion. Click to exit out of the pop-up window. Selecting the box for Don't show this again will bypass the pop-up window the next time the alignment wizard is started.
| The image uploaded in the manual alignment wizard must match the image passed to the spaceranger count pipeline for analysis. |
The Visium Manual Alignment wizard accepts the same types of image files as
Space Ranger. Loupe Browser 6.2 added support for .qptiff format. Refer to the input image recommendations page for more details.
| Image Format | |
|---|---|
| Brightfield Image | 24-bit color TIFF/BigTIFF or a JPEG |
| 16-bit grayscale TIFF/BigTIFF or a JPEG | |
| Fluorescence Image | 8 or 16-bit grayscale single, multi-page TIFF/BigTIFF | 8 or 16-bit grayscale multiple single-page TIFF/BigTIFF or a JPEG |
| 24-bit single colored image TIFF/BigTIFF or a JPEG | |
| CytAssist Image | 24-bit color TIFF |
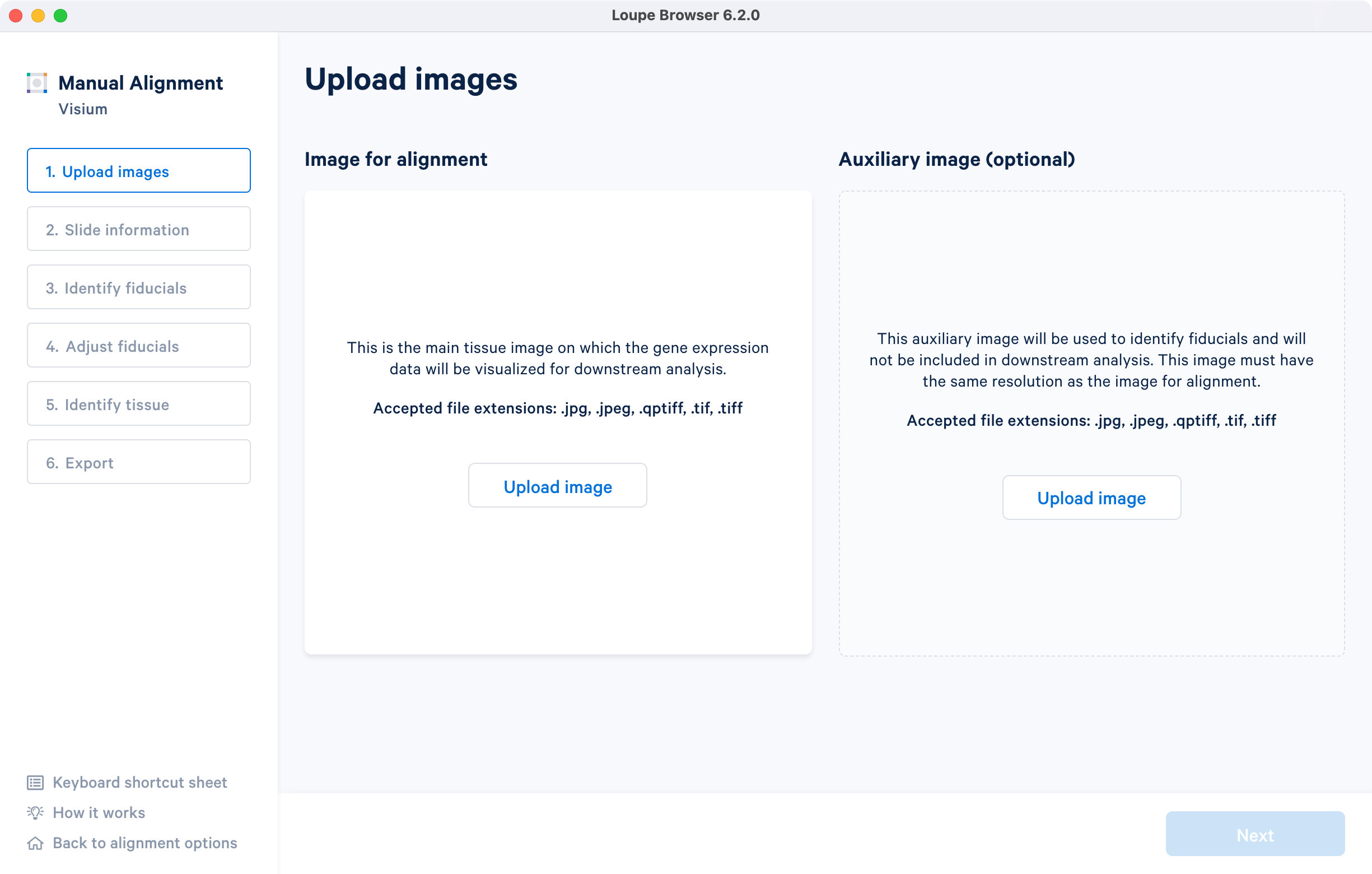
Clicking on under the Image for alignment panel opens a file dialog that filters the current filesystem for compatible image files. Selecting a file triggers a brief loading process. If the computer running the alignment wizard has insufficient RAM to efficiently load the image, the alignment wizard requires confirmation to continue. Note that clicking on the How it works on the lower left corner brings up the pop-up window.
Select the tab corresponding to the different starting image types.
The tutorial will use the following resources for the H&E-stained brightfield image.
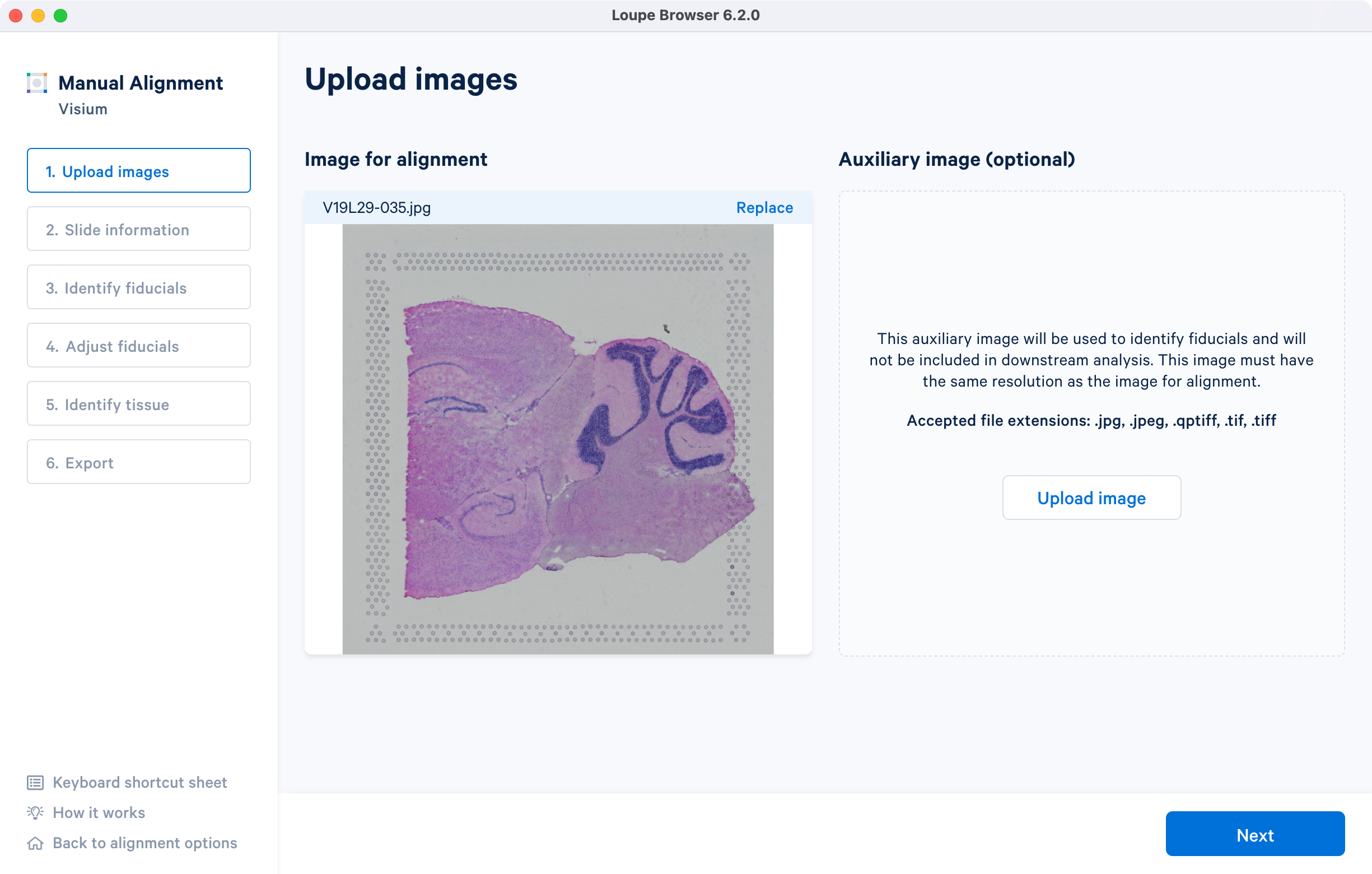
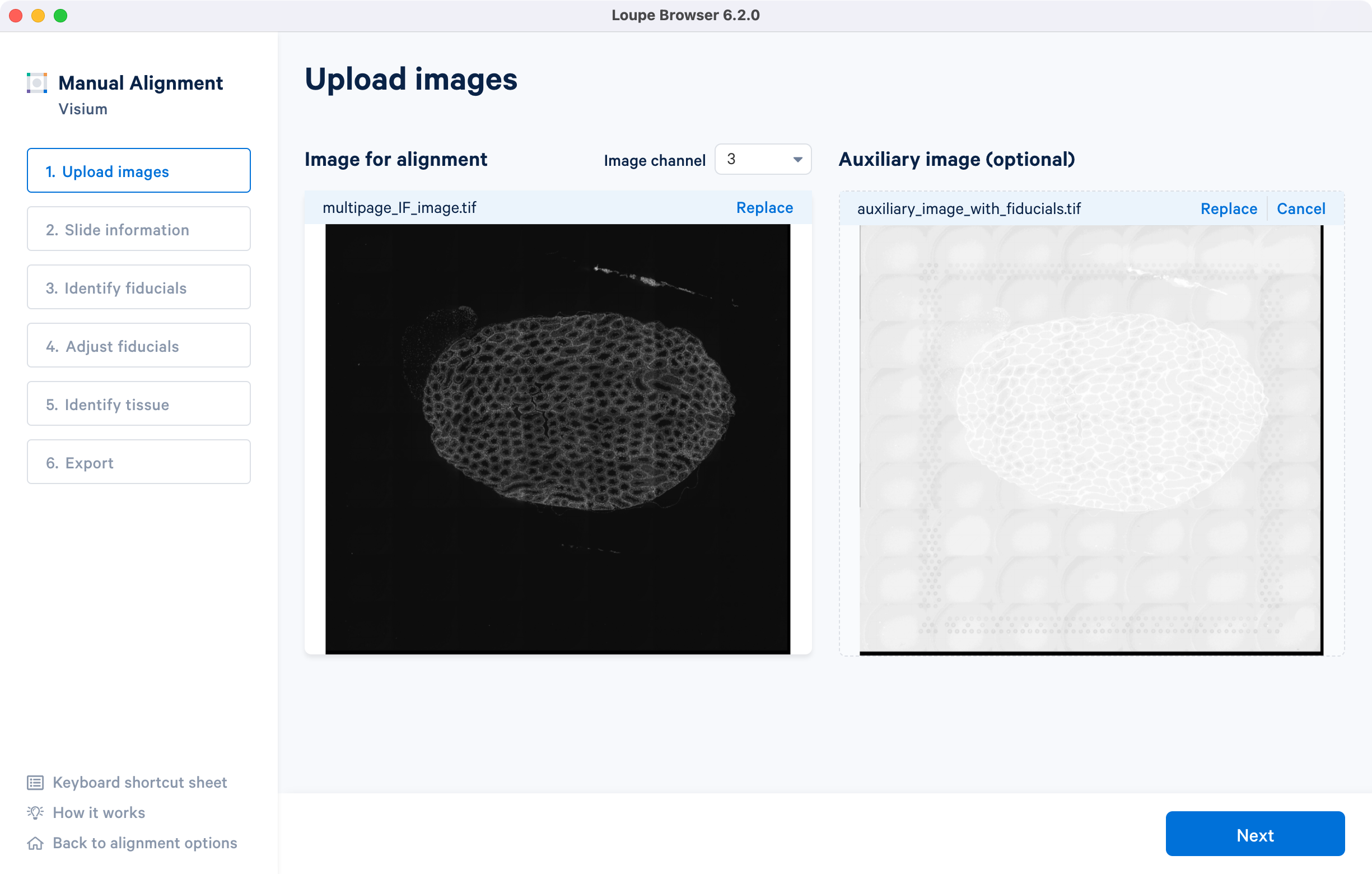
Once the image is loaded, preview image appear, as shown below. Successful upload of the image will make available for click. Use the Replace option located at the top right corner above image preview to change the loaded image.
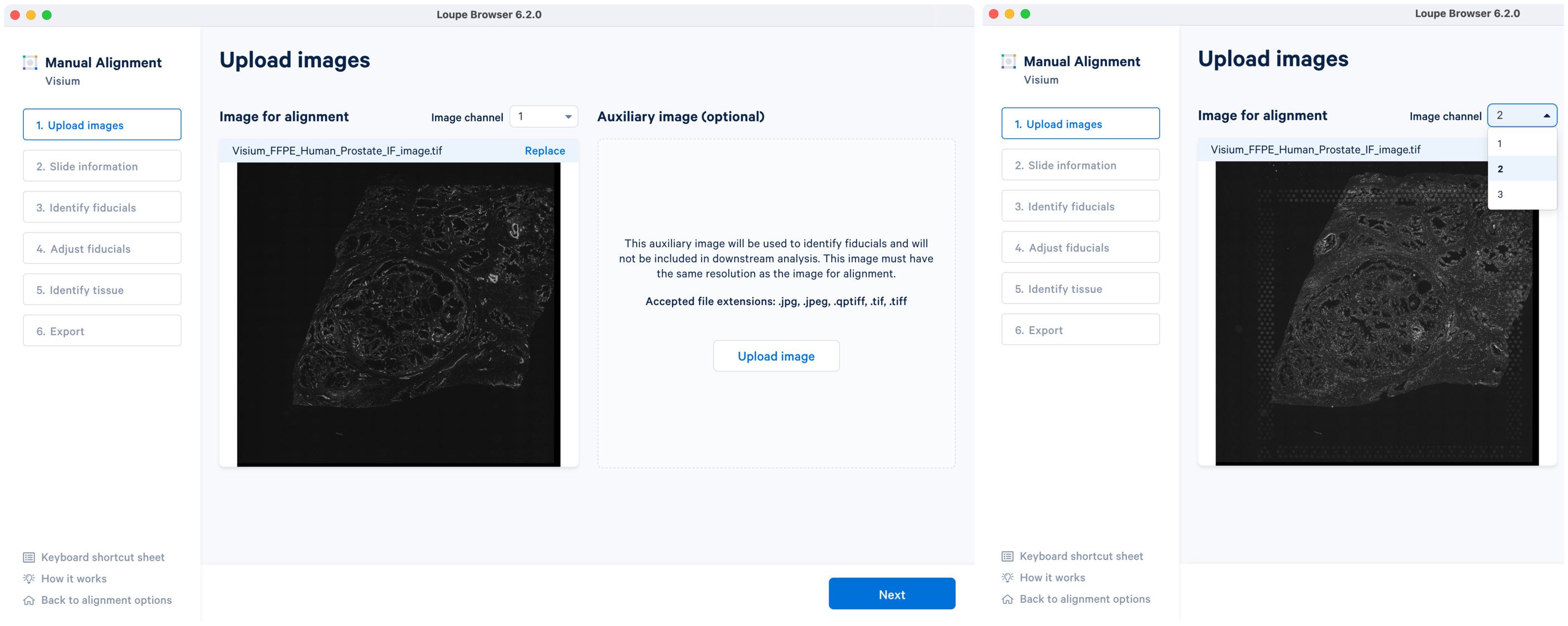
The tutorial will use multi-page fluorescence image included in human prostate FFPE public dataset.
Once the image is loaded, preview image appear, as shown below. Successful upload of the image will make available for click. Use the Replace option located at the top right corner above image preview to change the loaded image.
For multi-page fluorescence images, the fiducial markers and/or tissue may only be visible in a single page as shown for the tutorial dataset. Throughout the alignment wizard, use the Image channel dropdown to change the active page selection.
If an additional auxiliary overexposed image is being used for fiducial alignment, click on under the Auxiliary image (optional) panel and follow the same steps as above.
In case of multiple single page fluorescence images, there are three possible scenarios:
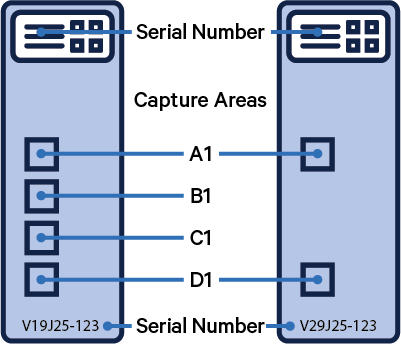
Visium slide serial number and capture areas are important parameters that are specified in this step.
 |
 |
There are three options for entering slide information in the Visium Manual Alignment wizard.
| Loupe Option | Internet Access | Details |
|---|---|---|
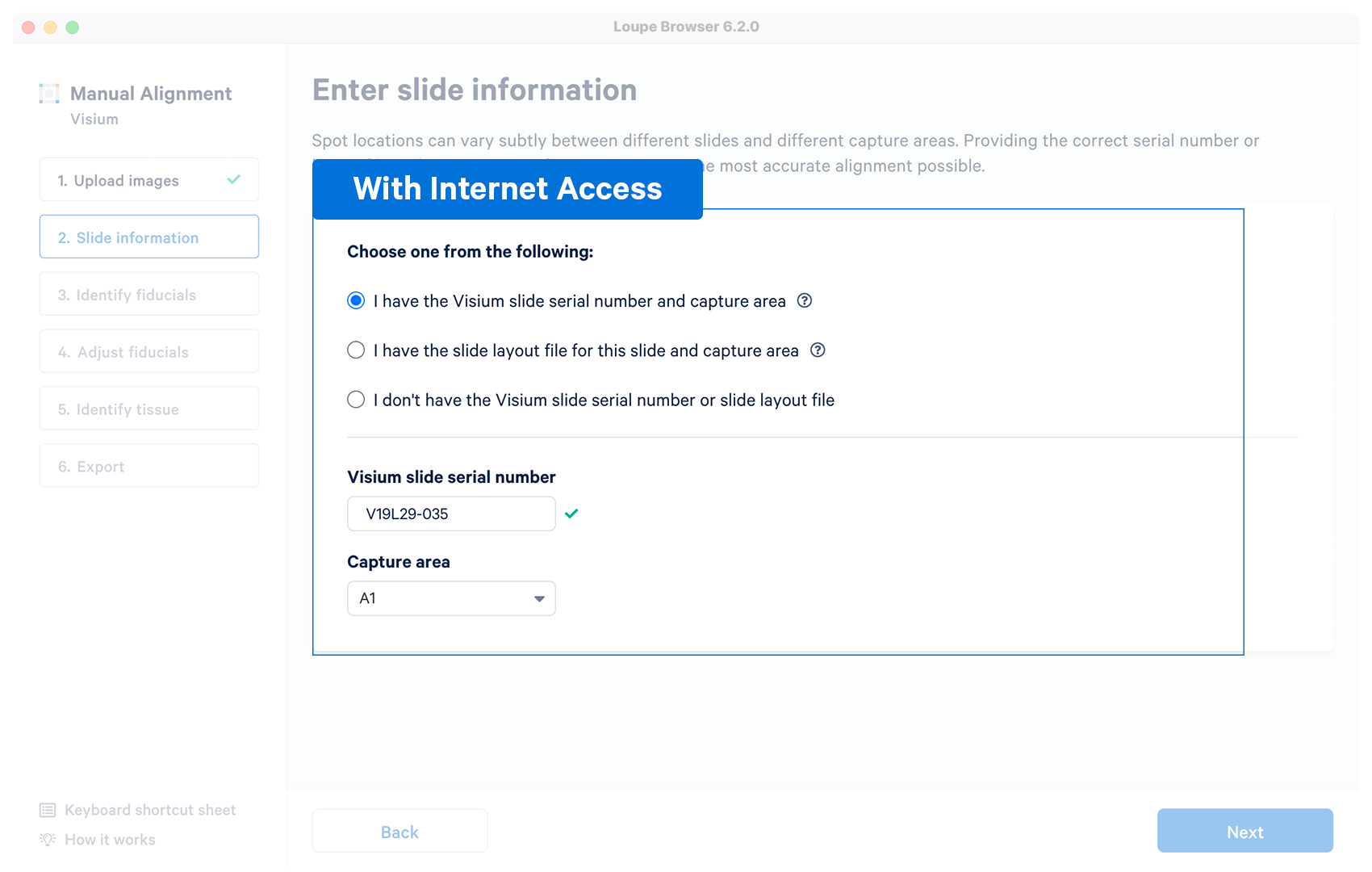 |
Yes | The alignment wizard will download the layout file associated with the supplied serial number. The alignment wizard shows a prompt if the slide number is invalid, or the machine does not have an internet connection. Capture area selector is visible after a successful entry. |
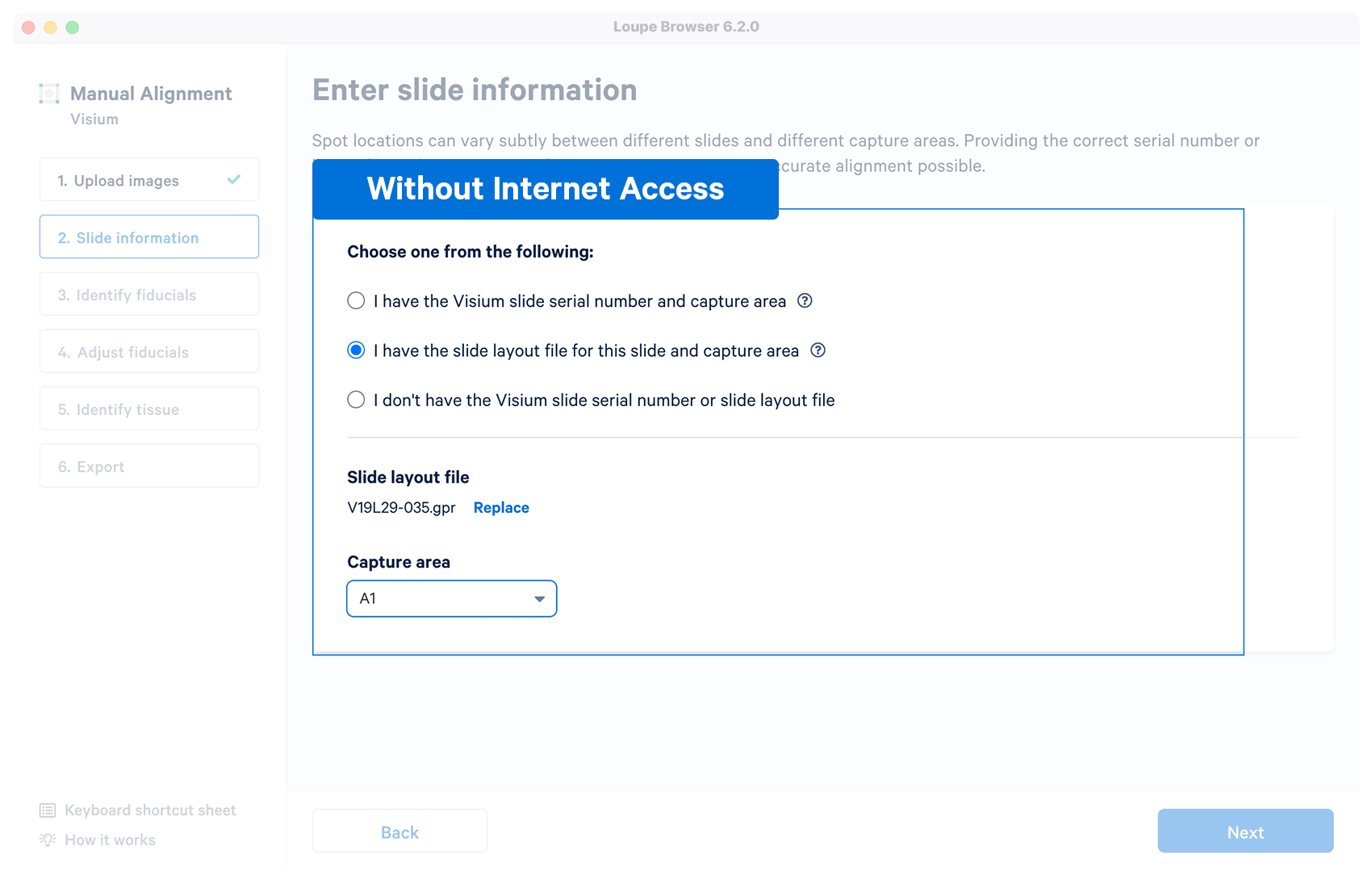 |
No | User downloads a layout file in gpr format for a Visium
slide before running the manual alignment. Once the file is loaded and validated, the capture area selector is visible. Download Slide Layout File |
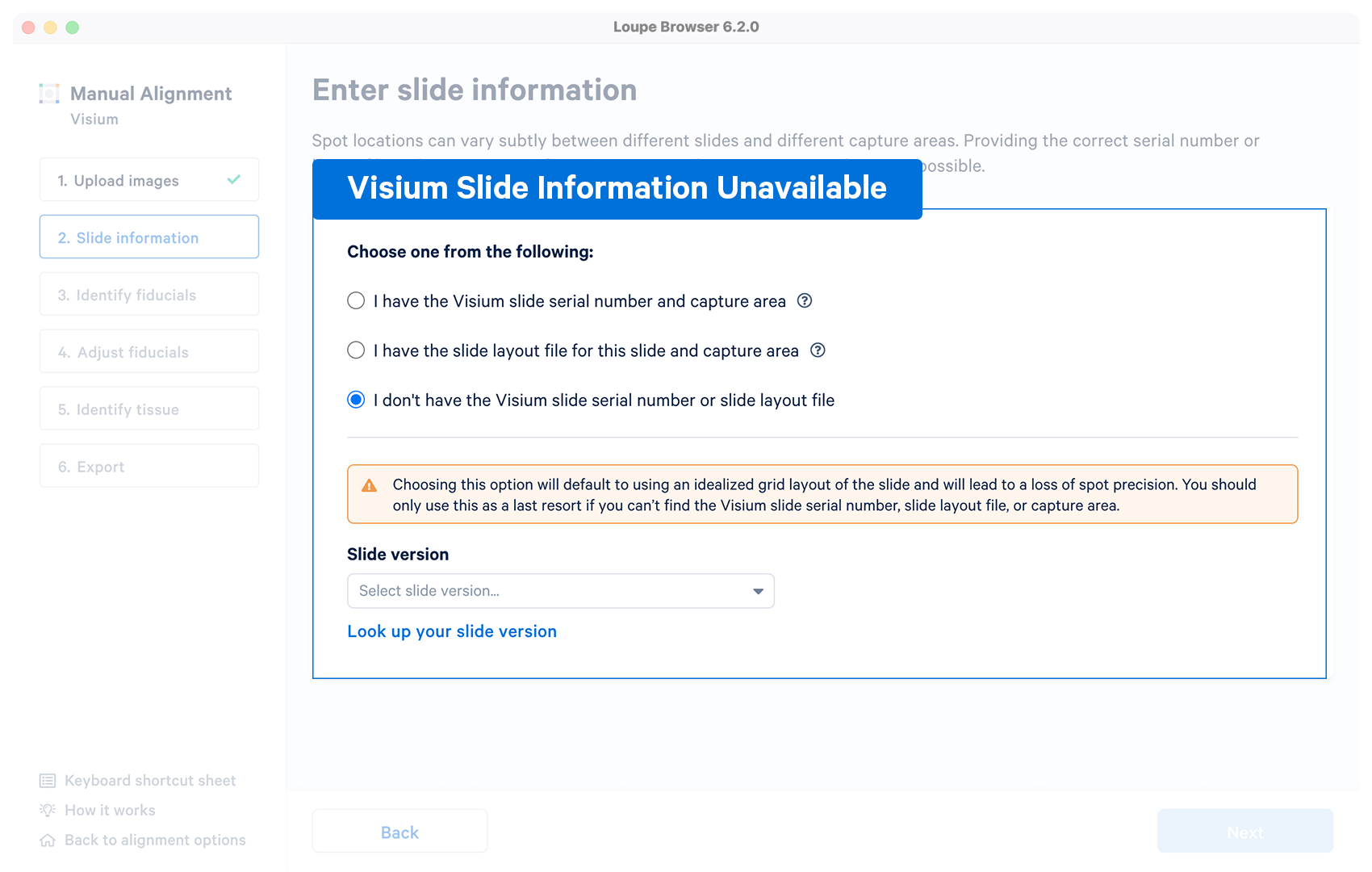 |
Optional | The alignment wizard uses an idealized grid layout in the alignment step corresponding to the slide version which can be choosen from the drop down menu. If unsure, click on Look up your slide version which will open a pop-up window with slide fiducial frames for reference. The typical per-spot difference between the default layout and a specific slide is within 5-15 microns, significantly smaller than the width of each spot. Refer to the Slide Parameters page for more information |
Select the tab corresponding to the different starting image types.
If you have internet access, enter the following slide serial number and select the capture area from the dropdown menu.
Appearance of a ✓ indicates successful retrieval. The slide number can be directly changed in the box.
To run the tutorial without internet access, you can download slide layout file in gpr format and upload it to the alignment wizard.
If you have internet access, enter the following slide serial number and select the capture area from the dropdown menu.
Appearance of a ✓ indicates successful retrieval. The slide number can be directly changed in the box.
To run the tutorial without internet access, you can download slide layout file in gpr format and upload it to the alignment wizard.
This step uses the spot location information from the image and the slide layout file loaded in the previous step to enable interactive manual alignment, using the Fiducial Markers as a visual guide. The spots are aligned with the image, first in a coarse-grained manner through identification of the centers of the Fiducial Markers then in a fine-grained manner by dragging to make small adjustments.
Loupe Browser 6.1 onwards new features and functionality were added that enhance the manual alignment experience. These are:
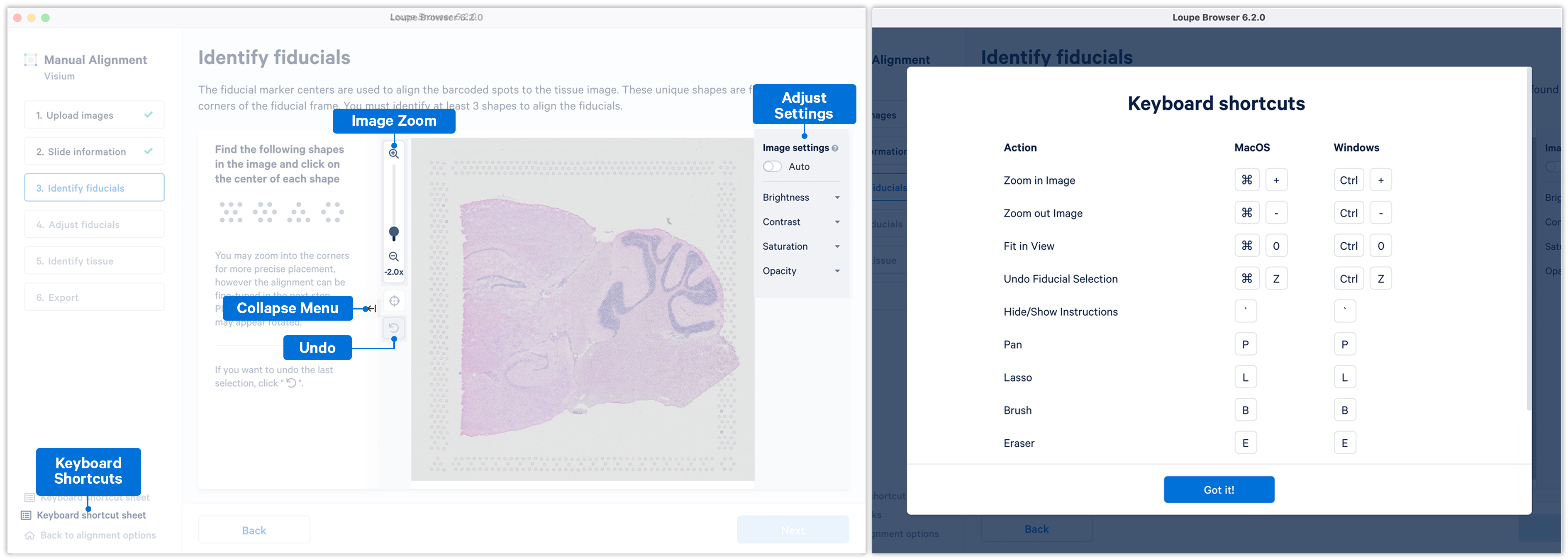
If the fiducial markers or tissue boundaries are dim, it is possible to adjust the brightness, contrast, saturation, and opacity of the image, which may make it easier to identify these features.
When set to on, the Auto toggle adjusts the brightness of the image across all the pixels with the goal to enhance the dark and bright pixels. It does so by mapping the original image brightness values [min, max] to a new range and rescaling it back to the full brightness spectrum [0, 255]. Note that the Auto toggle is independent of the manual image settings and can be used in combination with one or more of the other image settings.
| All the image settings apply evenly across all the pixels in a image as well as across all the channels in a multi-page TIFF. Saturation only applies to colored images. |
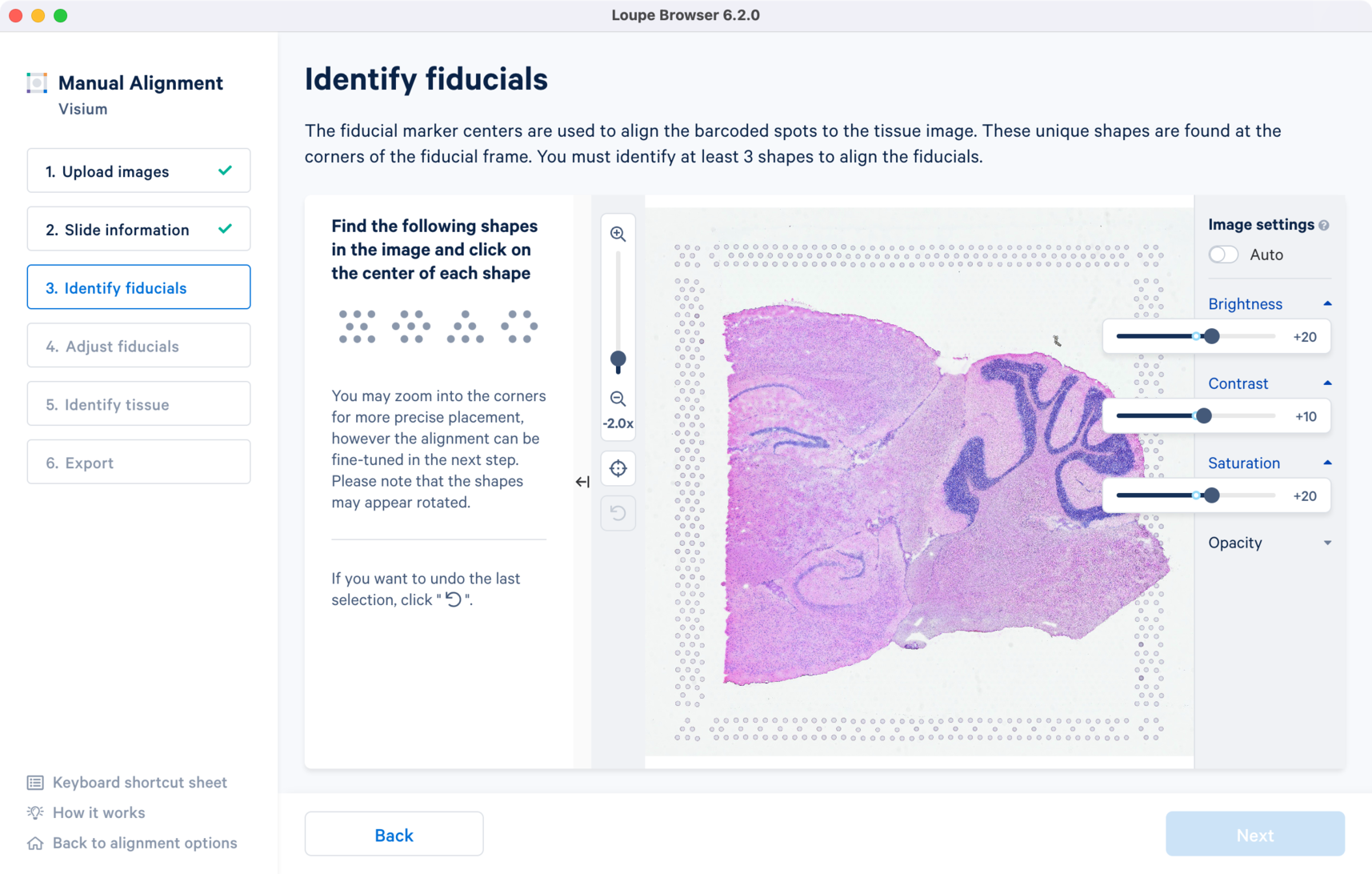
For multi-page fluorescence images or when using an auxiliary image, it is important to begin by selecting a page where fiducials are most clearly visible. Use the Image Channel dropdown to select the correct page. This may be an overexposed page in the same file that you plan to discard downstream in your analysis.
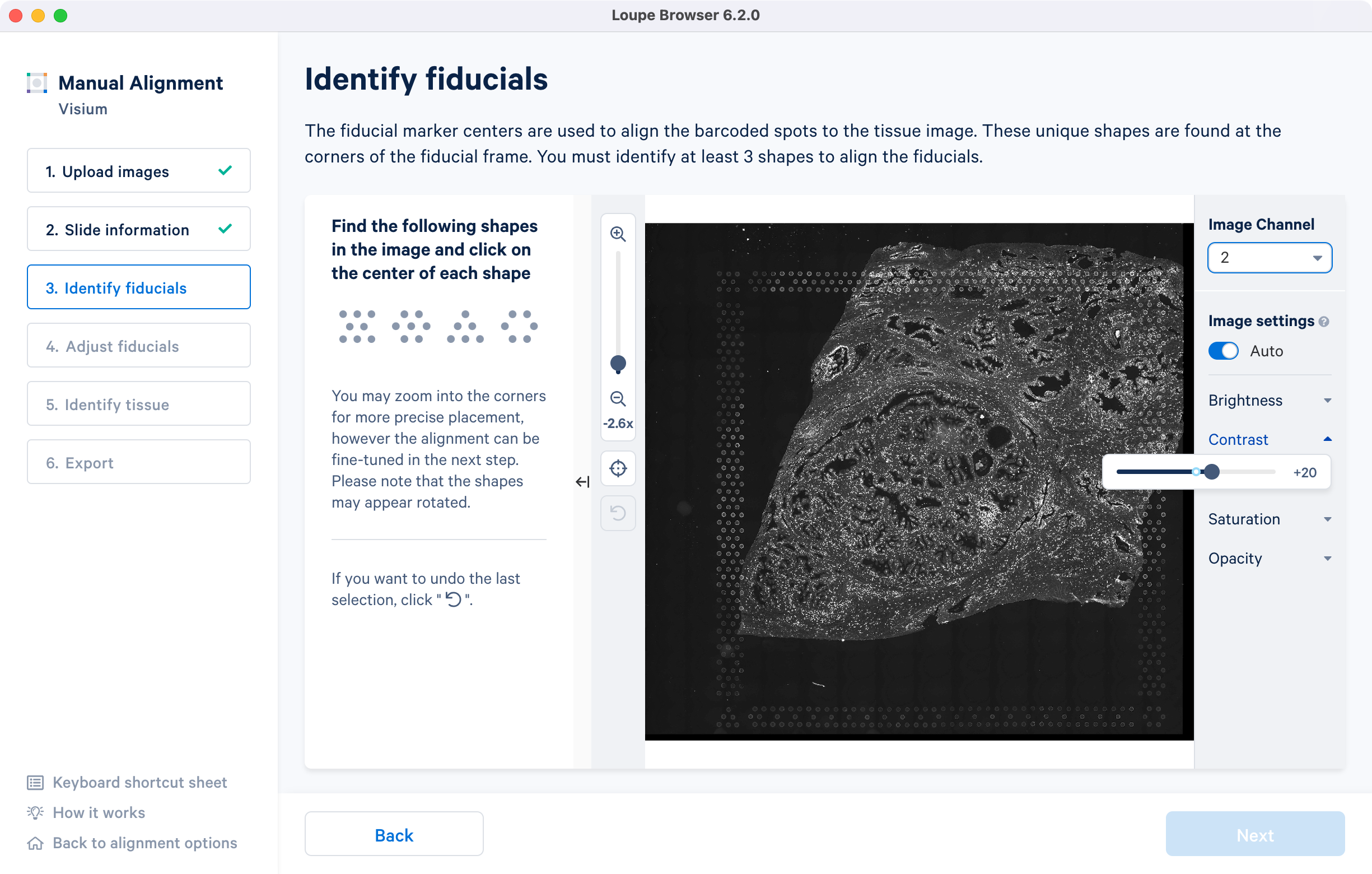
If an auxiliary overexposed image was loaded in the previous step, it will be accessible as an additional option in the Image Channel dropdown at the Identify fiducials step for fiducial selection. This image must have the same resolution as the first image loaded for alignment.
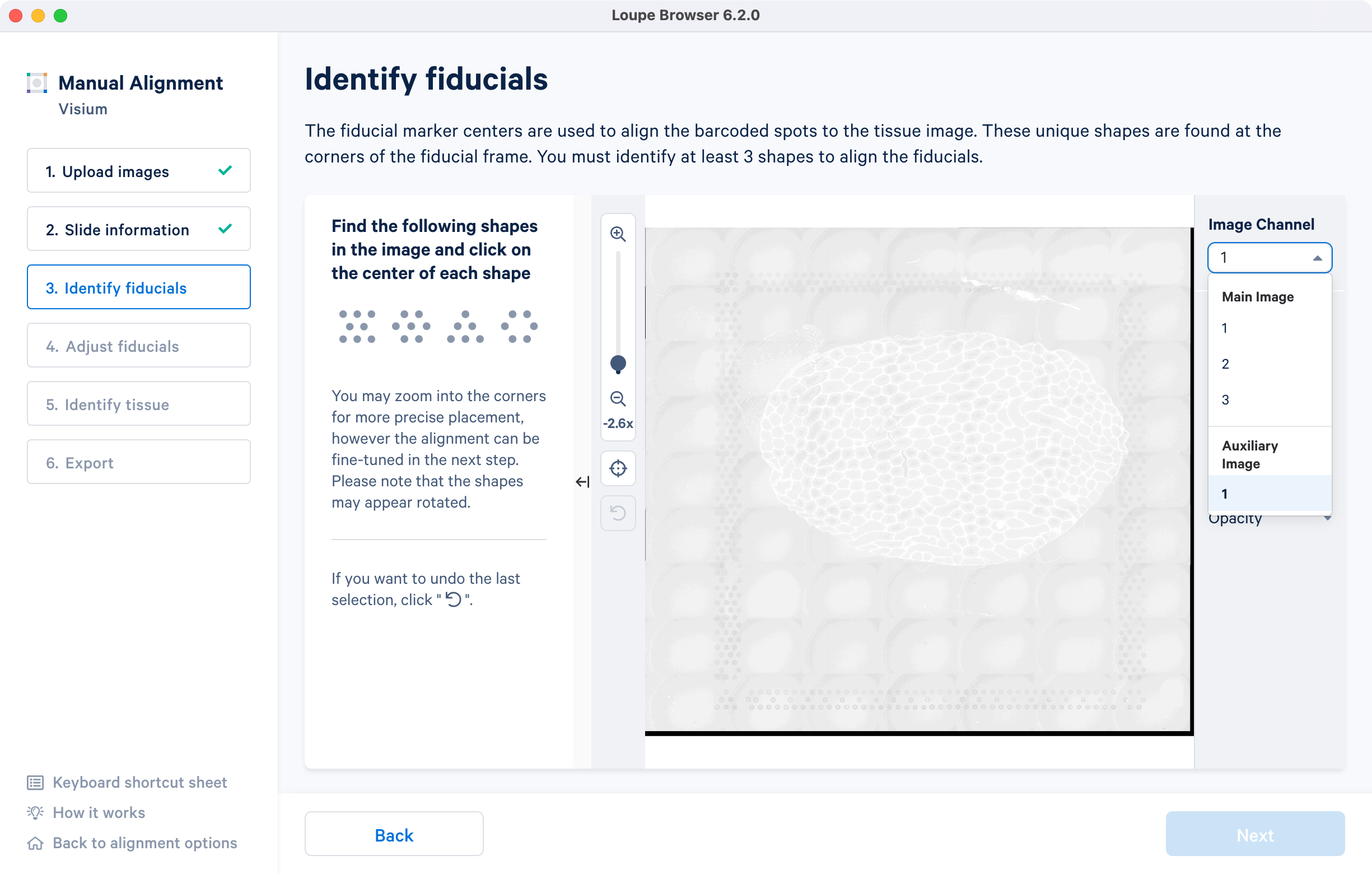
Visium slides have four unique fiducial corner shapes:
| Fiducial Markers | Shape |
|---|---|
| Hourglass | |
| Filled Hexagon | |
| Open Hexagon | |
| Triangle |
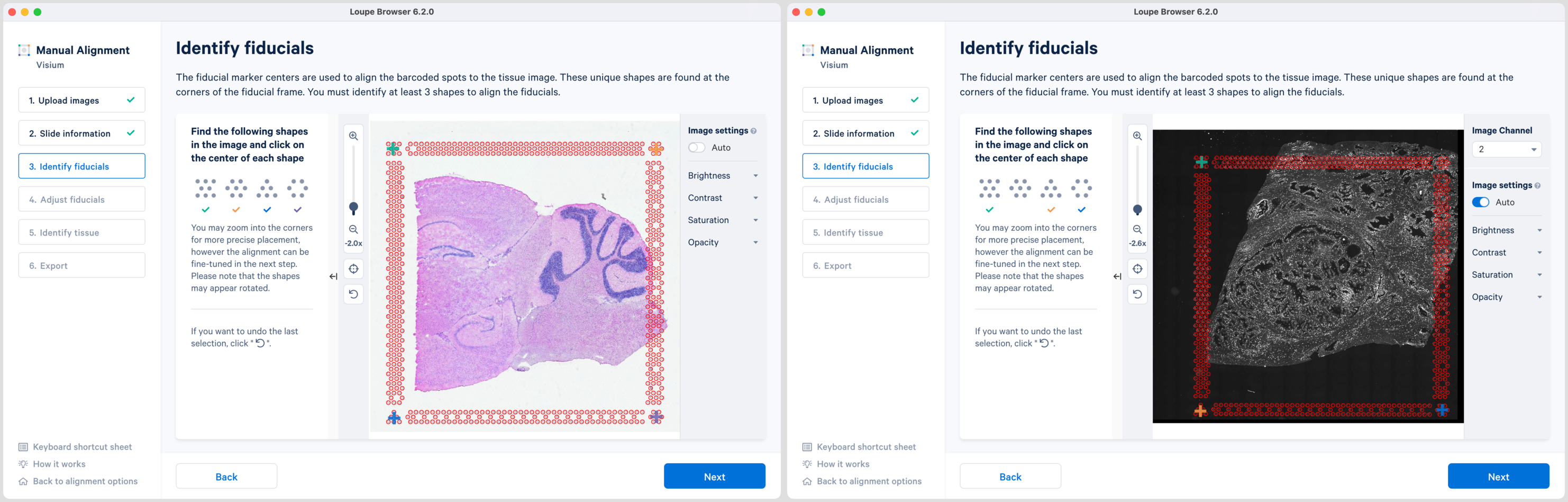
Alignment instructions are located on the left side of the page, and the alignment tool is on the right. It is important to follow the instructions carefully, as prompts change subtly at each step and it is critical to map each unique shape correctly.
Selection of each fiducial corner results in a colored + on the image and a matching ✓ on the instruction panel. Successful marking of at least three corners results in a red fiducial frame overlaid on top of the image. Click to revert selections at any time. Appearance of the red fiducial unlocks the to continue to step 4, Adjust fiducials.
| Fine-grained alignment by dragging the corner + can be performed both at Identify fiducials and Adjust fiducials steps. For better control over fine tuning it is recommended to mark all four fiducial markers if visible. |
While the placement of the red frame usually matches the fiducial frame on the image, in some instances fine-grained tuning may be required. In this step, you can achieve this dragging the corner + shapes to adjust the alignment of the red spot outlines to the fiducial frame in the image. Zoom in over the frame to identify misalignment and correct them by clicking and dragging the +. There are three options to zoom into the image using either the mouse wheel, zoom slider or the newly introduced keyboard shortcuts.
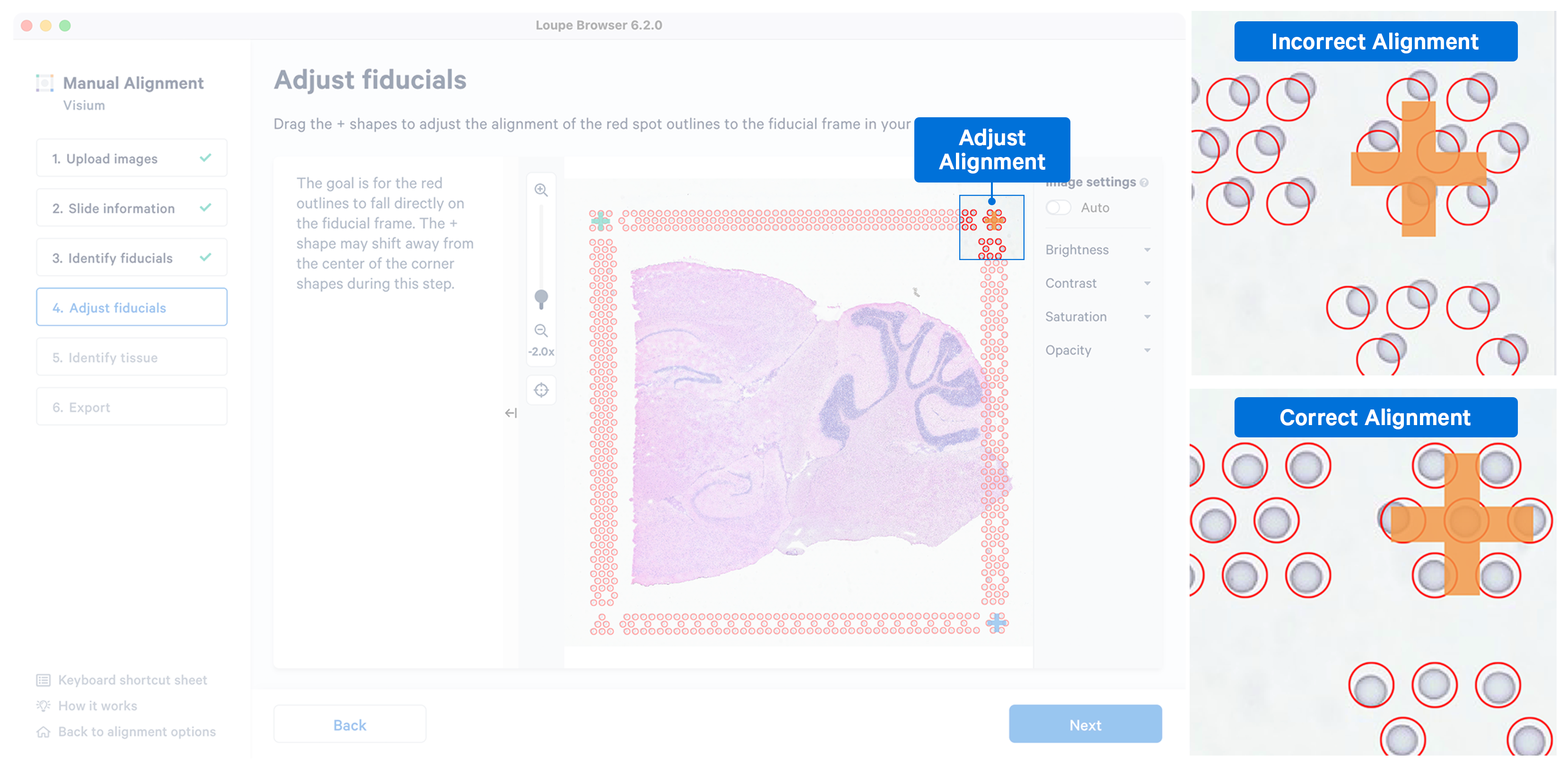
While fine-grained tuning is highly recommended to avoid any alignment errors, this step is optional and you can click to continue.
Accurate tissue boundary selection yields higher-quality data, lessens the likelihood of background-driven clusters, and provides accurate spot counts for fine-tuning sequencing parameters. When starting on the Identify Tissue page, the barcoded spots, with location inferred from the previous alignment step, are drawn on top of the image, as shown below. A toolbar to the left of the image provides a variety of options to perform both coarse-grained and fine-grained tissue selection.
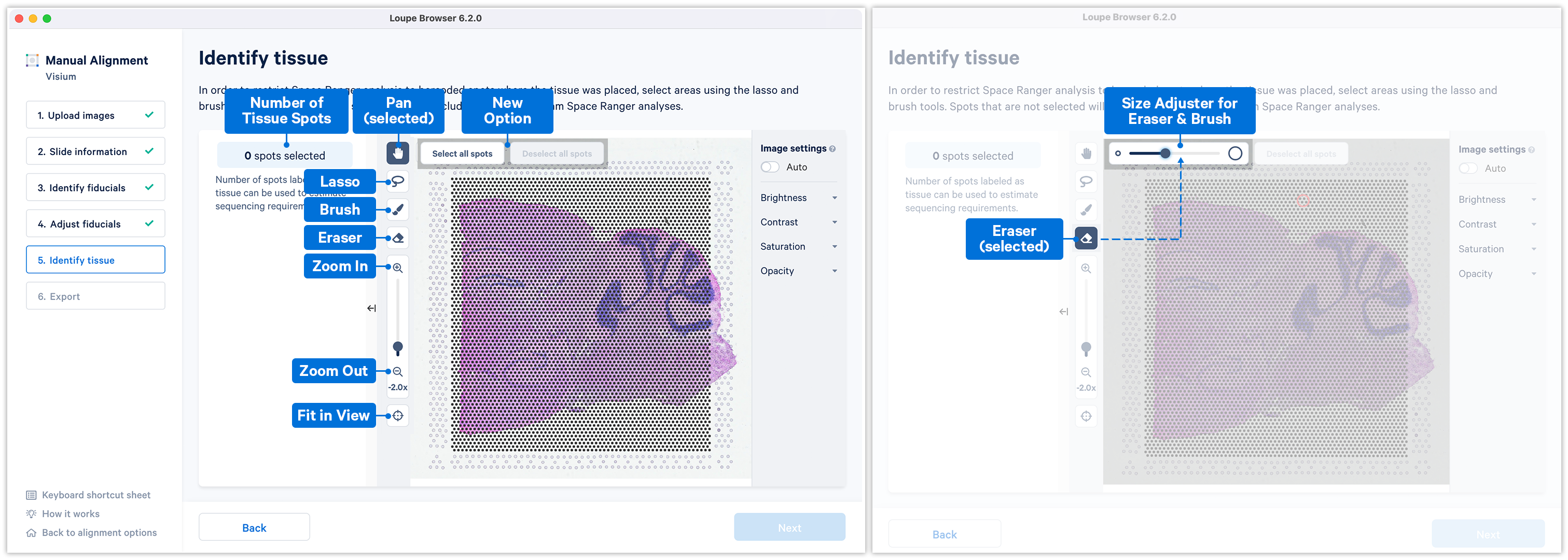
For multi-page fluorescence images, the page that contains the clearest tissue boundaries may be different from the page that most clearly shows the fiducial markers. The active page can be changed by clicking on the Image Channel dropdown under the tissue alignment instructions. For the tutorial dataset set the page to 3 for this step.
Key details for when using the tools:
There are two possible ways to select the tissue associated spots which appear green upon selection:
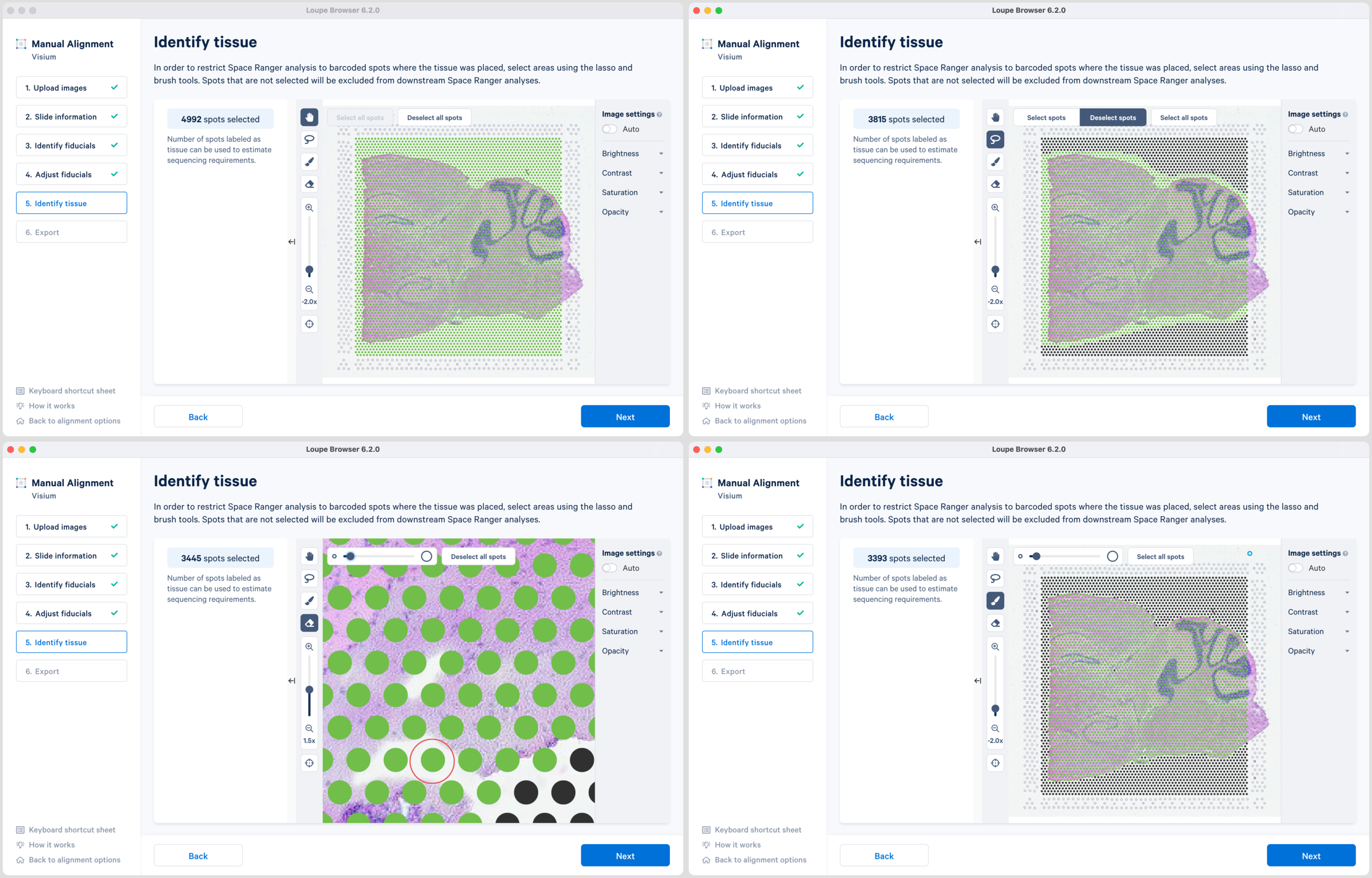
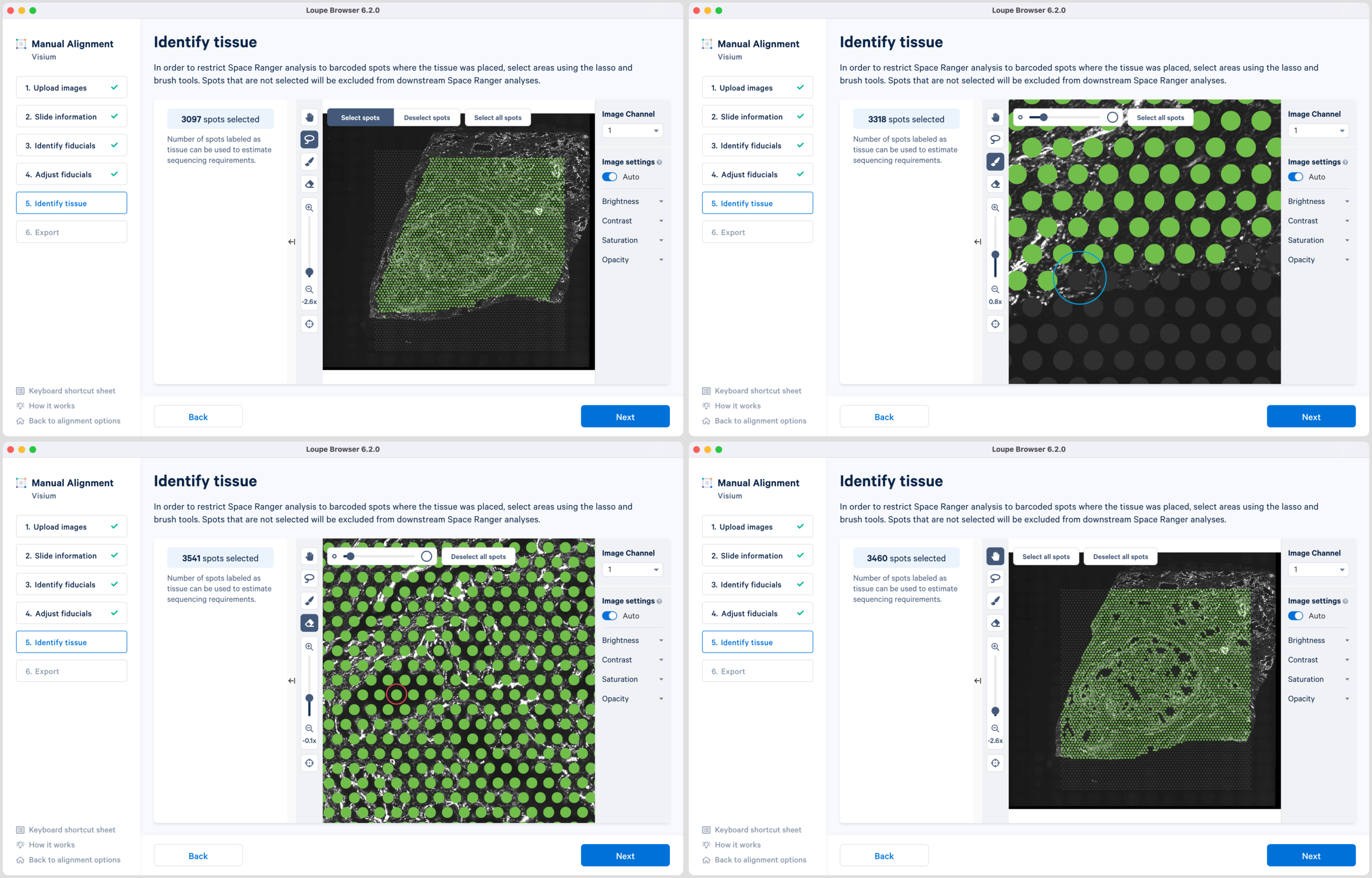
Note that you can return to previous steps in the alignment wizard to fine-tune, make adjustments, restart alignment, restart tissue detection, or enter different data. If alignment is satisfactory, click to continue to the last step in the manual alignment worklfow.
The final step shows the complete fiducial spot alignment and tissue selection. Click to generate a JSON file, which is named after the slide and capture area, if it was supplied in Slide information step.
| The JSON file generated after performing manual alignment in Loupe Browser contains important information about the spot pixel coordinates. This file is used in Space Ranger to override automatic fiducial and tissue detection. The internal format of this file is not fixed and modification of this file is not recommended. If you wish to modify the alignment, repeat the manual alignment process in Loupe Browser to generate a new JSON file. |
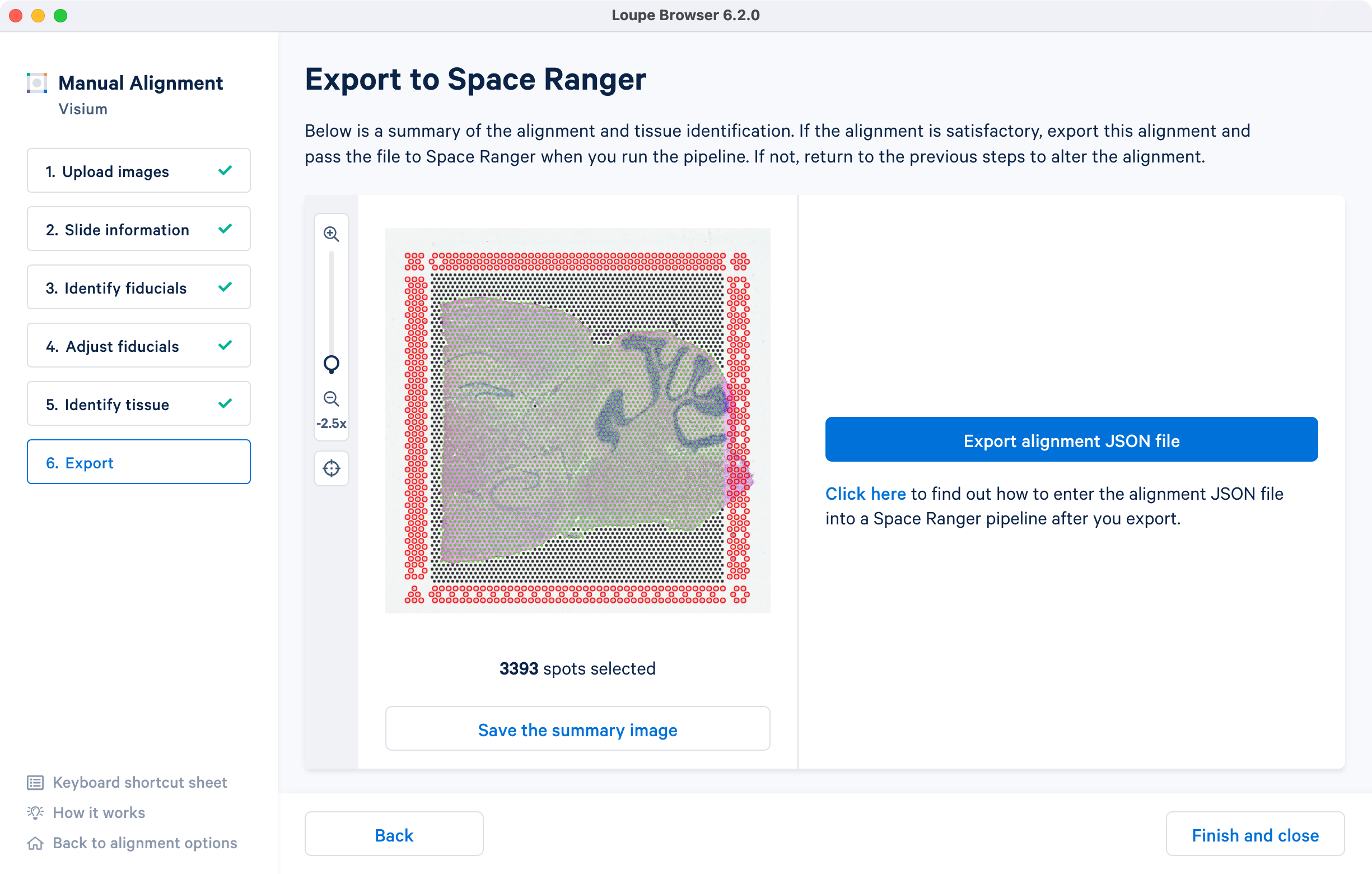
The image roughly mirrors the alignment preview shown in a Space Ranger web summary file. Use option for record-keeping purposes. Click here opens the Space Ranger count page in a new tab in a browser window.
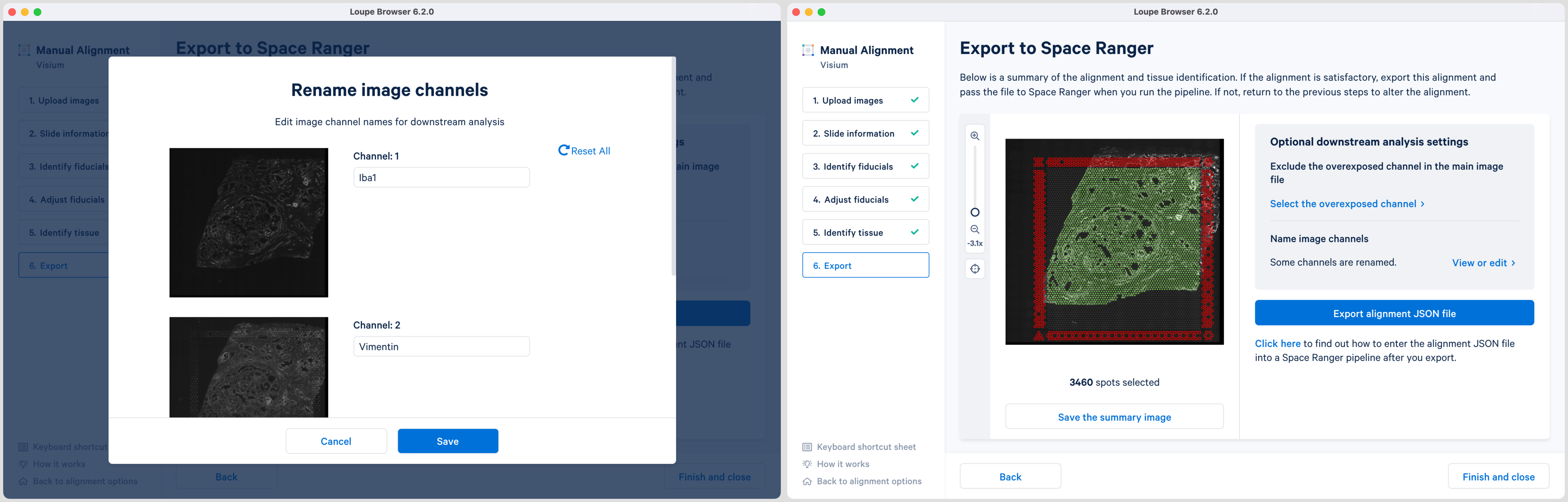
For multi-page fluorescence images where an overexposed capture was added as a page in the main image file, there is an option to exclude this page from downstream analysis. This is highly recommended to ensure that the tissue is clearly visible in the web summary and Loupe Browser analysis. Click Select the overexposed channel option and in the pop-up window select the page and click to exit. An auxiliary image will be excluded by default.
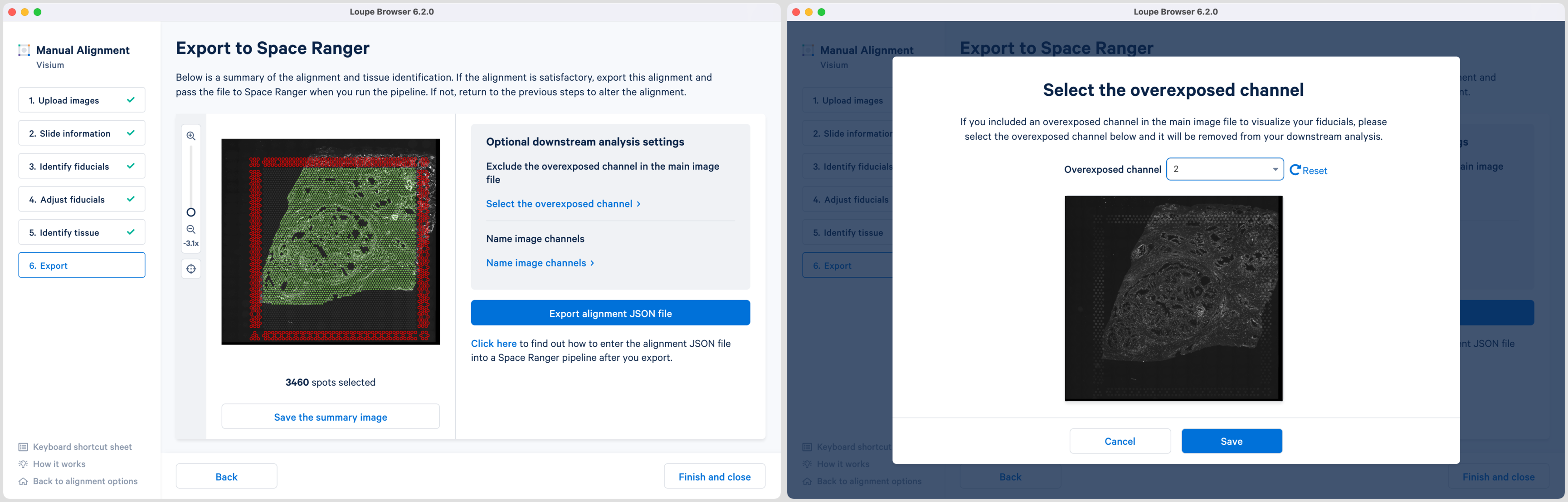
Loupe 6.2 also introduces the ability to name the fluorescence channels for multi-page TIFF images. The pop-up window is auto populated with the number of channels based on the input image. Click Name image channels option and in the pop-up window, type the name in the box under each Channel number and click to exit. By default, the channels are named as channel n where n denotes an integer number starting at 1. It is possible to name only a subset of the channels and the remainder will follow the default channel name scheme. All channels including the overexposed channel if applicable are shown in this view.
The export page is now updated to indicate channels were named. Click View or edit to view the changes in the pop-up window. If channel names are provided in Loupe Browser, then Space Ranger will use these for column names in barcode_fluorescence_intensity.csv file as well in the Loupe browser cloupe.cloupe output file.
The path to the generated JSON file is provided to spaceranger count pipeline via the --loupe-alignment argument. For more
information, consult the Manual Alignment instructions in the Space Ranger documentation. For workflows involving the CytAssist image, the Loupe alignment file can contain either both manual image registration and fiducial alignment information or only the manul fiducial alignment information. All possible combinations are listed in the Manual Alignment section of the spacerange count page for CytAssist.
Note: It is important to make the alignment file available to both spaceranger count and the computing environment where the pipeline is run.