Space Ranger2.0, printed on 03/23/2025
The pipeline output contains a subdirectory named spatial/, which stores spatial related files including quality control (QC) images. These QC images are useful to confirm accurate fiducial alignment and tissue detection. However, the user supplied original full resolution images are used for visualization in Loupe Browser.
Space Ranger 2.0 onwards supports the CytAssist enabled Gene Expression (GEX) analysis which introduces additional files in this folder.
Choose the tab to find the descriptions about the files in the folder corresponding to each workflow.
tissue_hires_image.png and tissue_lowres_image.png: These files are downsampled versions of the original,
full-resolution image provided by the user. Downsampling is accomplished by box filtering, which averages RGB values in patches of pixels in the full-resolution image to obtain an RGB value of one pixel in the downsampled image. Note that the downsampled images maintain the aspect ratio of the original image.
tissue_hires_image.png image has 2,000 pixels in its largest dimension.tissue_hires_image.png image is 4,000 pixels in its largest dimension.Irrespective of the slide type, tissue_lowres_image.png is always 600 pixels.
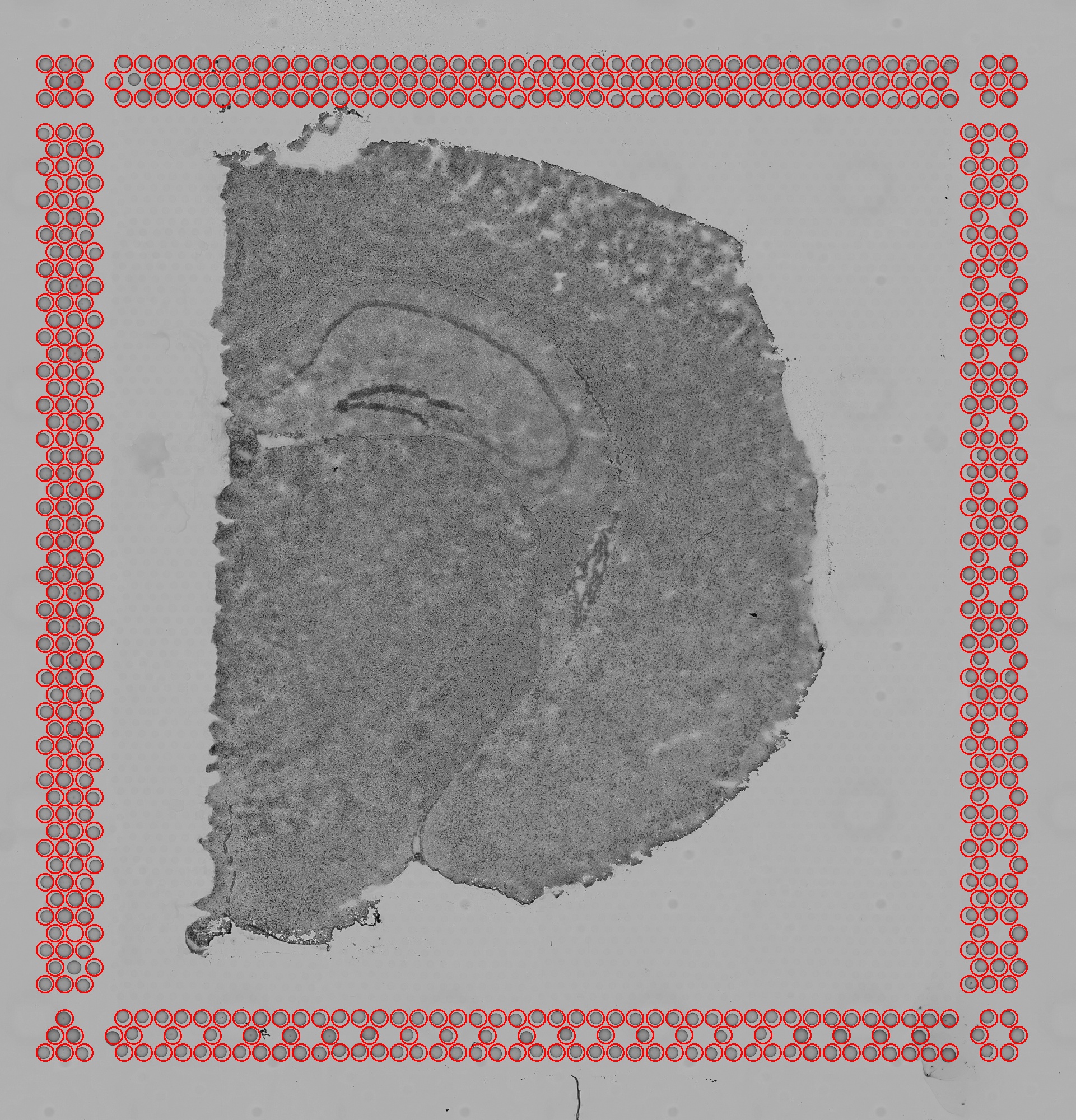
aligned_fiducials.jpg: This image has the dimensions of tissue_hires_image.png. Fiducial spots found by the
fiducial alignment algorithm are highlighted in red. This file is useful to verify that fiducial alignment was
successful.

detected_tissue_image.jpg: This image has the dimensions of the tissue_hires_image.png and shows the following:
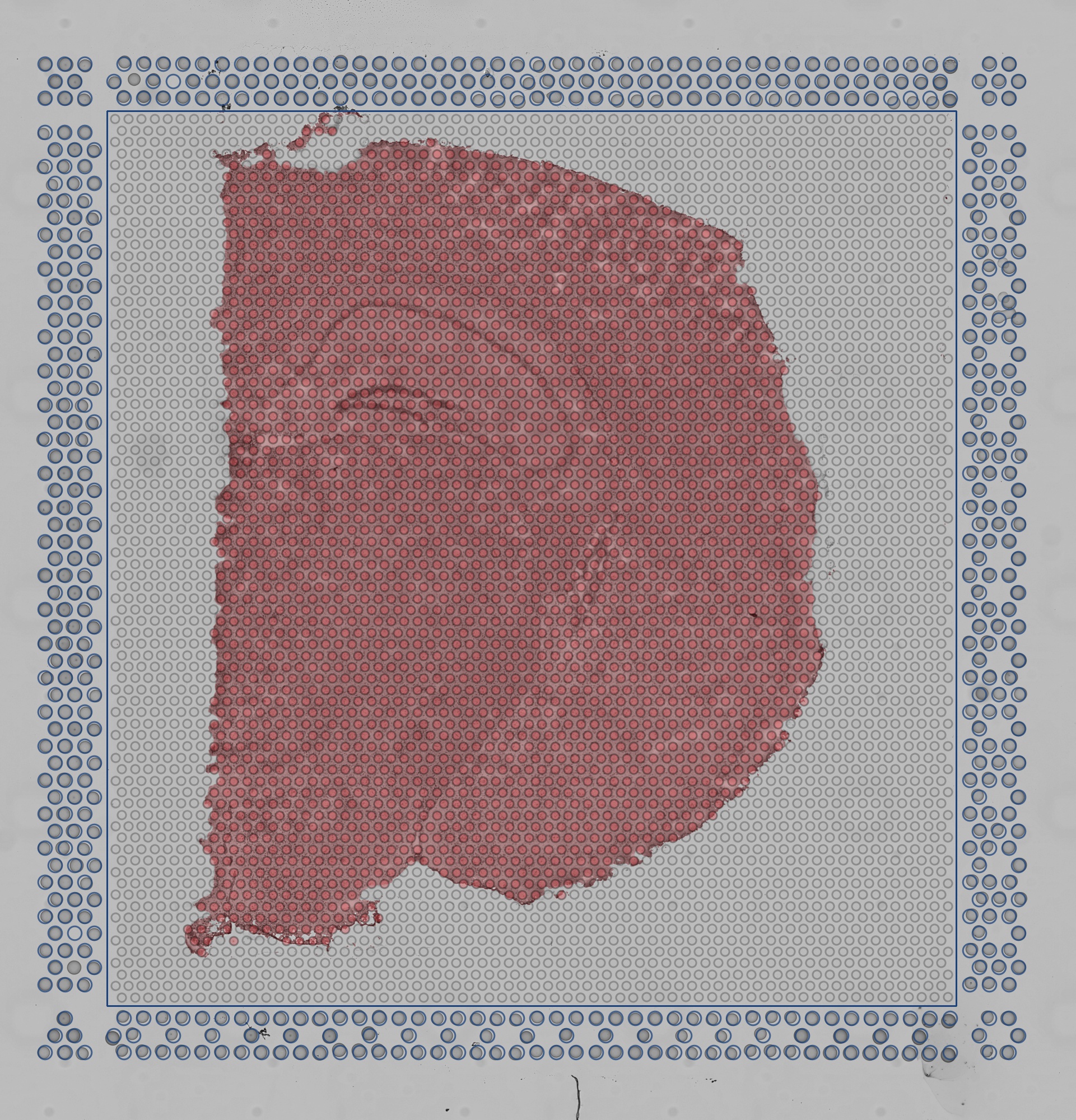

scalefactors_json.json: This file contains the following fields:
tissue_hires_scalef: A scaling factor that converts pixel positions in the original, full-resolution image to
pixel positions in tissue_hires_image.png.tissue_lowres_scalef: A scaling factor that converts pixel positions in the original, full-resolution image to
pixel positions in tissue_lowres_image.png.fiducial_diameter_fullres: The number of pixels that span the diameter of a fiducial spot in the original,
full-resolution image.spot_diameter_fullres: The number of pixels that span the diameter of a theoretical 65µm spot in the original, full-resolution image.$ cd /home/jdoe/runs/sampele345/outs/spatial $ cat scalefactors_json.json {"tissue_hires_scalef": 0.17011142, "tissue_lowres_scalef": 0.051033426", "fiducial_diameter_fullres": 144.4773339857, "spot_diameter_fullres": 89.43834961021503}
The example scalefactors_json.json values above are derived from the adult mouse brain dataset which includes an image with dimensions of 11291 x 11757. The scalefactors are calculated as
$$scalefactor = \frac{\text{target size}}{max(\text{original image width, original image height})}$$
Since the largest dimension for the tissue_hires_image.png is 2000 pixels, the tissue hires scalefactor will be $\text{tissue_hires_scalef} =\frac{2000}{11757} \approx 0.17011142$
Similarly for the tissue_lowres_image.png, the largest dimension is 600 pixels and the tissue lowres scalefactor is $\text{tissue_lowres_scalef} =\frac{600}{11757} \approx 0.051033426$
The spot diameter and fiducial diameter are estimated fractional pixels of the spot diameter in the original image. They are estimated based on the registration solution and the known size of the spots and fiducial frame and does not use prior knowledge of the image pixel sizes. In this example, the estimated pixel size was calculated from the spot diameter for the full resolution image. $\text{microns/pixels} =\frac{65}{89.43834961} \approx 0.7267575965$
tissue_positions.csv: This text file contains a table with rows that correspond to spots. From Space Ranger v2.0 onwards this file, which was previously named tissue_positions_list.csv, is renamed and includes a header column. Excluding the header column, the file has 4,992 rows for Visium slides with a 6.5 mm Capture Area and 14,336 rows for Visium slides with 11 mm Capture Area which is the number of spots in the spatial array. Columns correspond to the
following fields:
barcode: The sequence of the barcode associated to the spot.in_tissue: Binary, indicating if the spot falls inside (1) or outside (0) of tissue.array_row: For Visium slide (6.5 mm Capture Area), the row coordinate of the spot in the array from 0 to 77. The array has 78 rows. For Visium slides (11 mm Capture Area), the row coordinate of the spot range from 0 to 127 as the array has 128 rows.array_col: The column coordinate of the spot in the array. In order to express the "orange crate" arrangement of the spots, for Visium slides (6.5 mm Capture Area) this column index uses even numbers from 0 to 126 for even rows, and odd numbers from 1 to 127 for odd
rows with each row (even or odd) resulting in 64 spots. For Visium slides (11 mm Capture Area) this column index uses even numbers from 0 to 222 for even rows, and odd numbers from 1 to 223 for odd
rows with each row (even or odd) resulting in 111 spots.pxl_row_in_fullres: The row pixel coordinate of the center of the spot in the full resolution image.pxl_col_in_fullres: The column pixel coordinate of the center of the spot in the full resolution image.$ cd /home/jdoe/runs/sample345/outs/spatial $ column -s, -t < tissue_positions.csv | head -n 10 barcode in_tissue array_row array_col pxl_row_in_fullres pxl_col_in_fullres GTCACTTCCTTCTAGA-1 0 0 0 1832 11971 CACGGTCTCCTTACGA-1 0 0 2 1832 11833 ATAGCTGCGGATAAGA-1 0 0 4 1832 11695 GTCAGTATGTCCGGCG-1 0 0 6 1832 11556 ATGTACCAGTTACTCG-1 0 0 8 1831 11418 ACGCTCAGTGCACCGT-1 0 0 10 1831 11280 TCACTAACGTATAGTT-1 0 0 12 1831 11142 CGGTTAGGCCTGGACG-1 0 0 14 1831 11004 GATATCACCAGCATGG-1 0 0 16 1831 10866
The spots are arranged using a orange crate packing to maximize packing density. Further explanation of this arrangement can be found in this article.
Note that the tissue_positions.csv contains the same information about the barcode whitelist and their slide coordinates as that contained in the barcodes folder that comes bundled with Space Ranger (spaceranger-x.y.z/lib/python/cellranger/barcodes where x.y.z represent the version number).
spatial_enrichment.csv: File in CSV format that contains table of Moran's I values for each feature when specific conditions are met:
The Moran's I value can be anywhere between -1 (perfectly dispersed) to 1 (perfectly enriched) but in biological samples values significantly below 0 are unexpected. A p-value is provided, as well as an adjusted p-value which is corrected using the Benjamini-Hochberg method for multiple comparisons.
From Space Ranger 2.0 onwards, the spatial_enrichment.csv file includes a Feature Type column the value for which is Gene Expression for single-library analysis.
# Gene Expression output $ cd /home/jdoe/runs/sampele345/outs/spatial $ column -s, -t < spatial_enrichment.csv | less -#2 -S Feature ID Feature Name Feature Type I P value Adjusted p value Feature Counts in Spots Under Tissue Median Normalized Average Counts Barcodes Detected per Feature ENSMUSG00000053310 Nrgn Gene Expression 0.849230495400066 0.0 0.0 140702 42.22927672729207 2645 ENSMUSG00000070570 Slc17a7 Gene Expression 0.8228828454463434 0.0 0.0 49919 15.927820652112793 2435 ENSMUSG00000046447 Camk2n1 Gene Expression 0.8112121614961633 0.0 0.0 126518 44.058800706638436 2698 ENSMUSG00000021948 Prkcd Gene Expression 0.8093848987347875 0.0 0.0 11675 4.552770117948635 1430 ENSMUSG00000061808 Ttr Gene Expression 0.8088005899881938 0.0 0.0 348686 76.73215387483816 2674 ENSMUSG00000041607 Mbp Gene Expression 0.7965984861055619 0.0 0.0 258769 112.84701279639039 2700 ENSMUSG00000035383 Pmch Gene Expression 0.7847484395395488 0.0 0.0 67814 17.005396313242215 1718 ENSMUSG00000024985 Tcf7l2 Gene Expression 0.7769492292430982 0.0 0.0 6546 2.564212465278306 1282 ENSMUSG00000032532 Cck Gene Expression 0.7605729556575779 0.0 0.0 67935 21.68537496001758 2585
barcode_fluorescence_intensity.csv: File in CSV format that is generated for any input image specified by --darkimage. This file contains the statistics about the fluorescence intensity which is calculated for every spot and every fluorescence channel in the input image.
If the channels were given custom names in the Loupe manual alignment session that produced the alignment JSON file, then the CSV header will use the given channel names. By default, channels are named channel1, channel2, etc. This file can be used for secondary analysis in R/Python and can also be combined with the tissue_positions.csv file.
$ cd /home/jdoe/runs/sampele345/outs/spatial $ column -s, -t < barcode_fluorescence_intensity.csv | less -#2 -S barcode in_tissue channel1_mean channel1_stdev channel2_mean channel2_stdev CGAGGATATTCAGAGC-1 0 4254.328631875882 59.72596599289198 2370.7574047954868 23.587614594004506 TCTGGTACTAATGCGG-1 0 4242.12976022567 61.31391702047626 2348.9830747531737 22.960436841214605 AGGTGACCATGACGCT-1 0 4237.420310296192 62.954555749425516 2330.277856135402 18.66240280182469 CAGAGGTGTACGTAGC-1 0 4219.421720733428 60.38810661890725 2323.2045133991537 20.876943016829802 TAGCTAGAAGATACAT-1 0 4202.056417489422 62.2343918192054 2309.320169252468 18.883665883691563 AGCACTGACCGGCTTA-1 0 4185.466854724965 61.848950071485 2300.589562764457 17.98861741514344 ATAACCTAAGGCGTCA-1 0 4149.37658674189 58.4482514520289 2292.1255289139635 19.824612785901802 TAGAACCTCAGTTAGA-1 0 4156.802538787024 64.86614573399514 2298.7390691114247 19.838300058320733 CCGACATAAGTATCAA-1 0 4194.483779971792 62.19636612986138 2313.277856135402 20.56714424032058 GACTACCAACTGGTCC-1 0 4204.746121297602 62.91221526144468 2328.5232722143865 54.82377031240396
CytAssist enabled GEX analysis offers flexibility of using the CytAssist instrument captured image along with a microscope image of the same tissue section on a standard glass slide. The images of the spatial/ folder change depending on if a microscope image input was provided or not.
| The full resolution original image used for visualization corresponds to the microscope image (if one was provided) or to the CytAssist image. |
aligned_tissue_image.jpg: This image is generated when both CytAssist and microscope image inputs are provided. It captures the results of the image registration by merging the registered images in a checkerboard pattern with adjacent squares alternating between the two images. Successful image registration can be confirmed by assessing the continuity of tissue structures (e.g. tissue edges and interior tissue morphology). Note that dark background fluorescence microscope images are inverted to have a white background. An interactive version of these images is available in the Image panel of web_summary.html.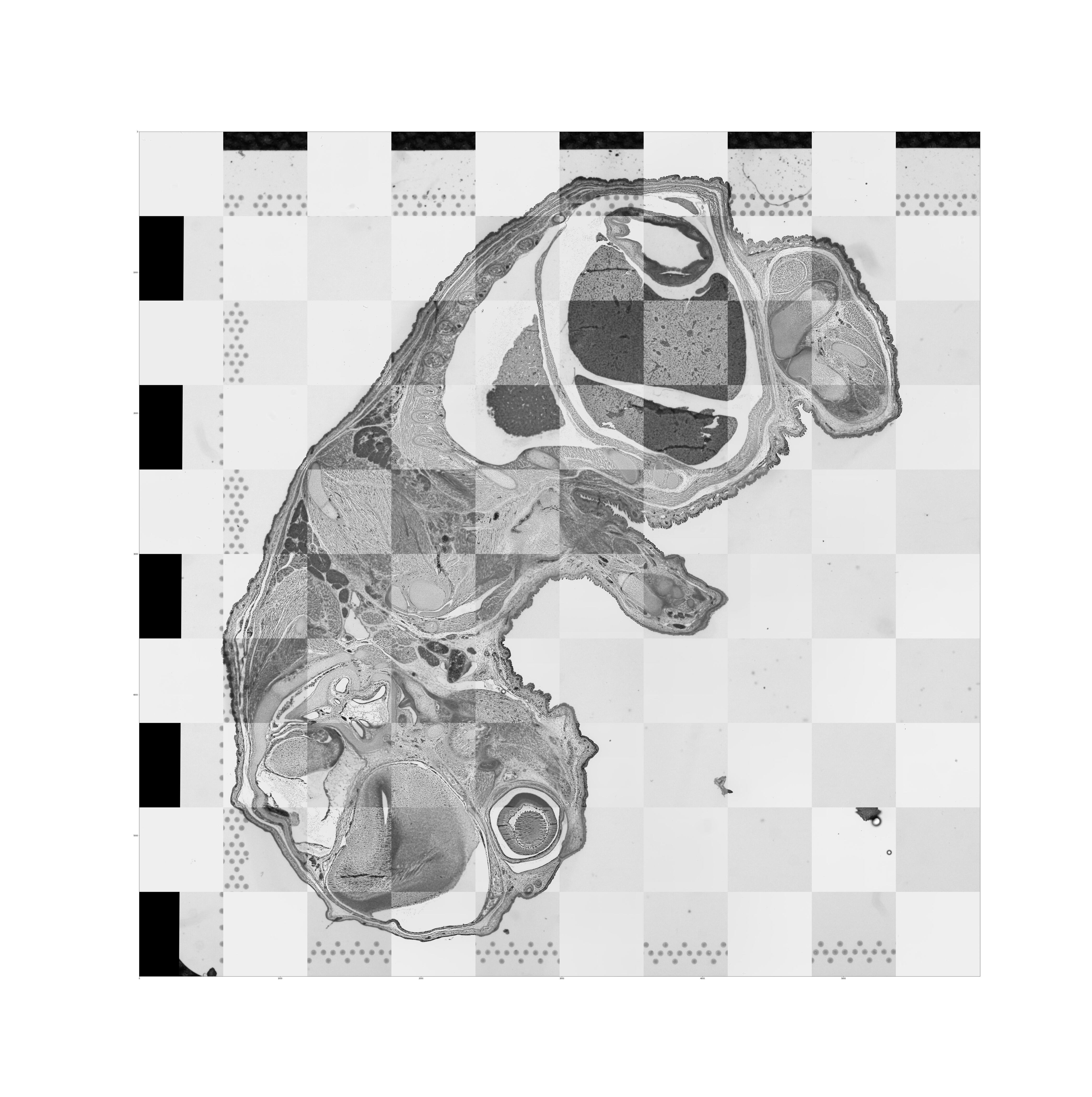
cytassist_image.tiff: Copy of the original CytAssist image input which is a eosin-stained brightfield image in TIFF format that contains the fiducial frame and has a fixed resolution of 3,000 pixels in both dimensions.
tissue_hires_image.png and tissue_lowres_image.png: These files are downsampled versions of the original, full-resolution image inputs provided by the user. Downsampling is accomplished by box filtering, which averages RGB values in patches of pixels in the full-resolution image to obtain an RGB value of one pixel in the downsampled image. Note that the downsampled images maintain the aspect ratio of the original image.
tissue_hires_image.png image has 2,000 pixels in its largest dimension. The tissue_lowres_image.png is set at 600 pixels. Both images corresponds to the CytAssist image.tissue_hires_image.png image corresponds to the microscope image and has 2,000 pixels in its largest dimension. The tissue_lowres_image.png is set at 600 pixels and also corresponds to the microscope image.aligned_fiducials.jpg: This image is always the CytAssist image with a dimension of 3,000 pixels in each dimension. Fiducial spots detected by the fiducial alignment algorithm are highlighted in red. The aligned_fiducials.jpeg contains information from the red channel since the fiducial frame is visible in it, and thus, the lack of tissue section contrast is expected. This file is useful to verify that fiducial alignment was successful.
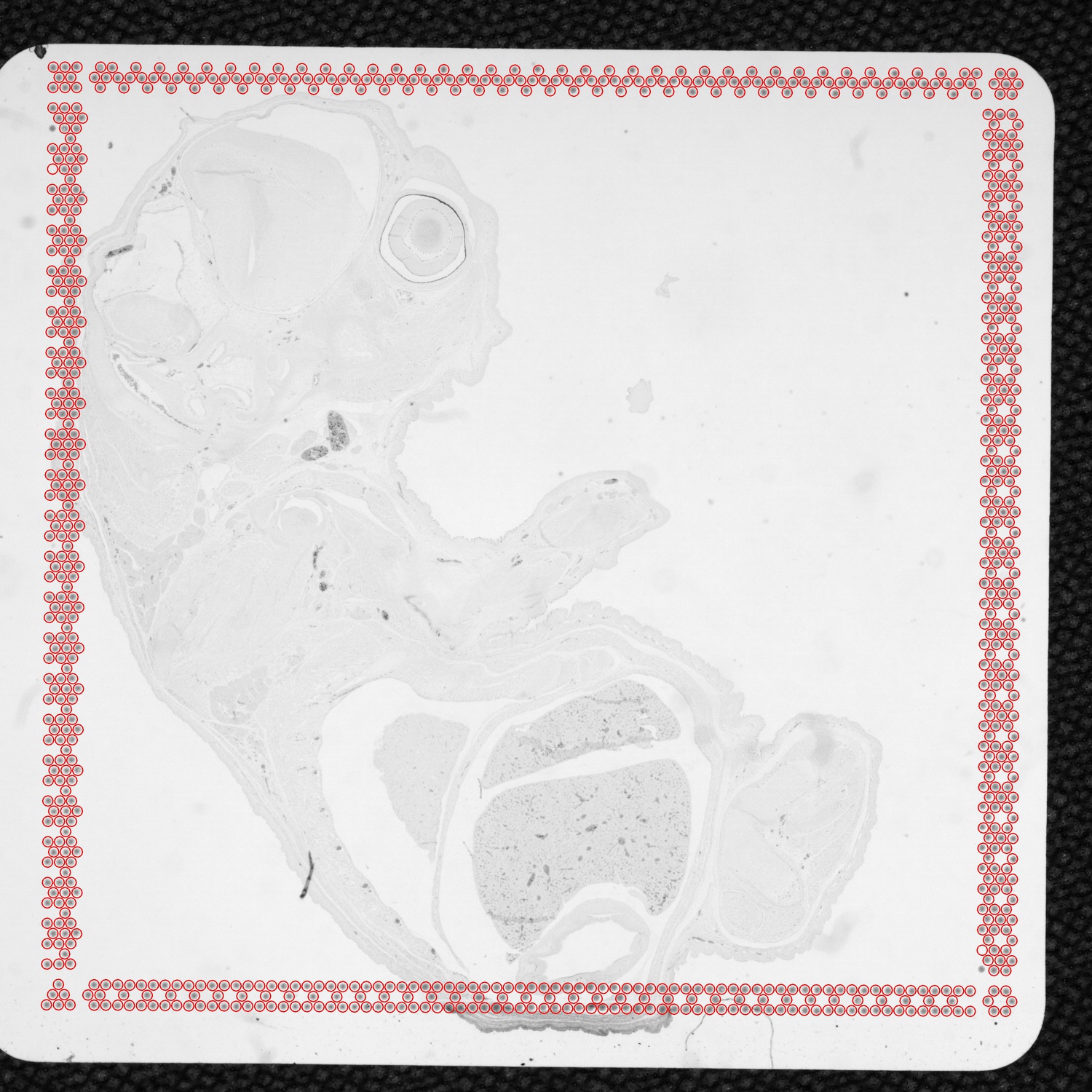
detected_tissue_image.jpg: This image is always the CytAssist image with a dimension of 3,000 pixels in each dimension and shows the following:
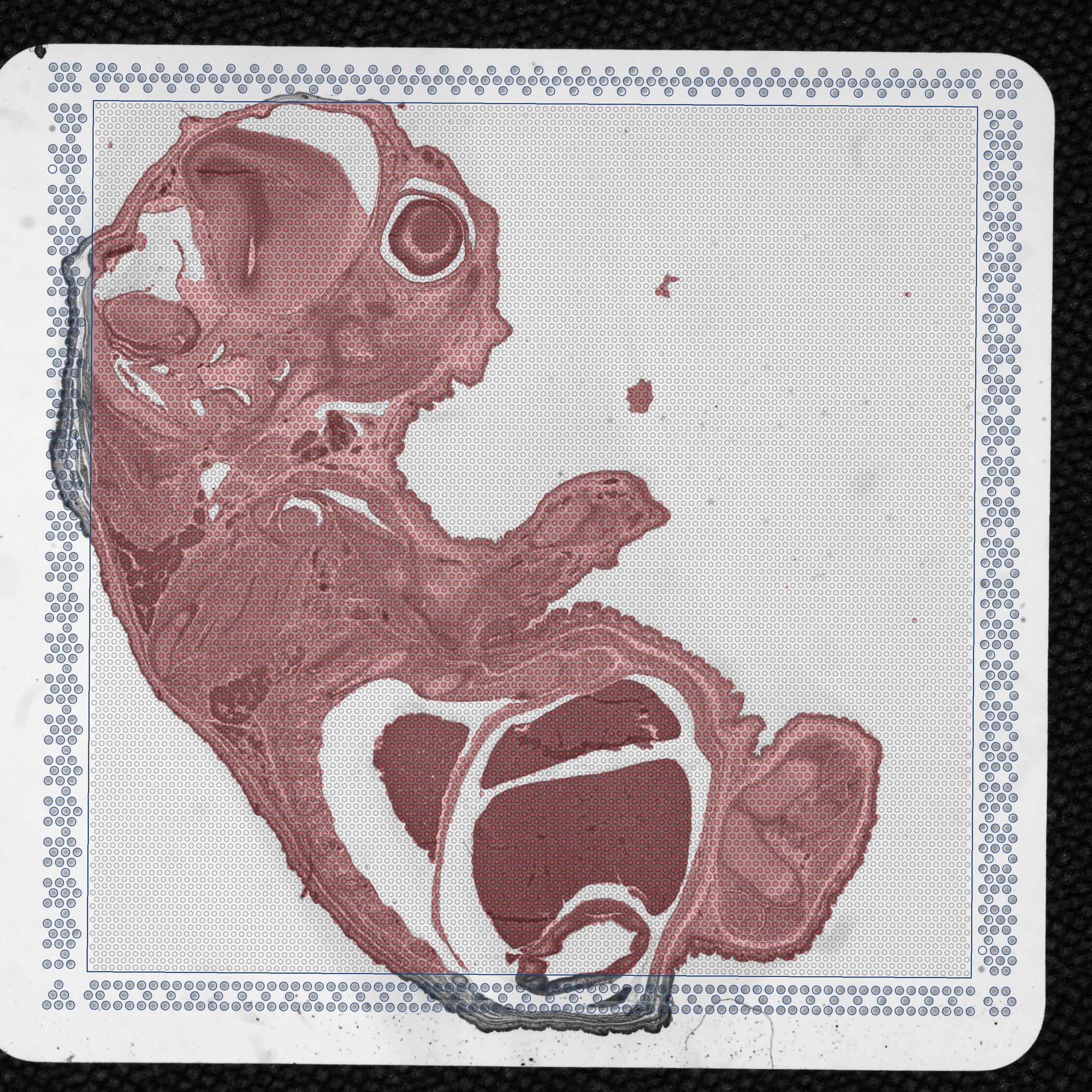
scalefactors_json.json: This file contains the following fields:
regist_target_img_scalef: A scaling factor that converts pixel positions in the original, full-resolution micrsocope image to the downsampled version used in image registration.tissue_hires_scalef: A scaling factor that converts pixel positions in the original, full-resolution image to
pixel positions in tissue_hires_image.png.tissue_lowres_scalef: A scaling factor that converts pixel positions in the original, full-resolution image to
pixel positions in tissue_lowres_image.png.fiducial_diameter_fullres: The number of pixels that span the diameter of a fiducial spot in the original,
full-resolution image.spot_diameter_fullres: The number of pixels that span the diameter of a theoretical 65µm spot in the original, full-resolution image.$ cd /home/jdoe/runs/sampele345/outs/spatial $ cat scalefactors_json.json {"regist_target_img_scalef": 0.45004502, "tissue_hires_scalef": 0.150015, "tissue_lowres_scalef": 0.045004502, "fiducial_diameter_fullres": 146.4305300684766, "spot_diameter_fullres": 97.62035337898439}
Assuming a microscope image with dimensions of 13332 x 11757. The scalefactor is calculated as $$scalefactor = \frac{\text{target size}}{max(\text{original image width, original image height})}$$
For image registration, the microscope image is downscaled to 6000 pixels in the largest dimension, so the scalefactor will be $\text{regist_target_img_scalef} =\frac{6000}{13332} \approx 0.4500450$
The regist_target_img_scalef will have a value of 1 if the microscope image resolution is < 6000 pixels as the original image input will be used in image registration.
Since the largest dimension for the tissue_hires_image.png is 2000 pixels, the scalefactor will be $\text{tissue_hires_scalef} =\frac{2000}{13332} \approx 0.150015$
Similarly for the tissue_lowres_image.png, the largest dimension is 600 pixels and the scalefactor is $\text{tissue_lowres_scalef} =\frac{600}{11757} \approx 0.04500450$
The spot diameter and fiducial diameter are estimated fractional pixels of the spot diameter in the original CytAssist image. It is estimated based on the registration solution and the known size of the spots and fiducial frame and does not use prior knowledge of the image pixel sizes. For this example, the estimated pixel size was calculated from the spot diameter for the full resolution image $\text{microns/pixels} =\frac{65}{97.62035337898439} \approx 0.66584475$
tissue_positions.csv : This text file contains a table with rows that correspond to spots. From Space Ranger v2.0 onwards, this file, which was previously named tissue_positions_list.csv, is renamed and includes a header column. Excluding the header column, the file has 4,992 rows for Visium slides with a 6.5 mm Capture Area and 14,336 rows for Visium slides with an 11 mm Capture Area, which is the number of spots in the spatial array. Columns correspond to the
following fields:
barcode: The sequence of the barcode associated to the spot.in_tissue: Binary, indicating if the spot falls inside (1) or outside (0) of tissue.array_row: For Visium slide (6.5 mm Capture Area), the row coordinate of the spot in the array from 0 to 77. The array has 78 rows. For Visium slides (11 mm Capture Area), the row coordinate of the spot range from 0 to 127 as the array has 128 rows.array_col: The column coordinate of the spot in the array. In order to express the orange crate arrangement of the spots, for Visium slides (6.5 mm Capture Area) this column index uses even numbers from 0 to 126 for even rows, and odd numbers from 1 to 127 for odd
rows with each row (even or odd) resulting in 64 spots. For Visium slides (11 mm Capture Area) this column index uses even numbers from 0 to 222 for even rows, and odd numbers from 1 to 223 for odd
rows with each row (even or odd) resulting in 111 spots.pxl_row_in_fullres: The row pixel coordinate of the center of the spot in the full resolution image.pxl_col_in_fullres: The column pixel coordinate of the center of the spot in the full resolution image.$ cd /home/jdoe/runs/sample345/outs/spatial $ column -s, -t < tissue_positions.csv | head -n 10 barcode in_tissue array_row array_col pxl_row_in_fullres pxl_col_in_fullres GTCACTTCCTTCTAGA-1 0 0 0 1832 11971 CACGGTCTCCTTACGA-1 0 0 2 1832 11833 ATAGCTGCGGATAAGA-1 0 0 4 1832 11695 GTCAGTATGTCCGGCG-1 0 0 6 1832 11556 ATGTACCAGTTACTCG-1 0 0 8 1831 11418 ACGCTCAGTGCACCGT-1 0 0 10 1831 11280 TCACTAACGTATAGTT-1 0 0 12 1831 11142 CGGTTAGGCCTGGACG-1 0 0 14 1831 11004 GATATCACCAGCATGG-1 0 0 16 1831 10866
The spots are arranged using a orange crate packing to maximize packing density. Further explanation of this arrangement can be found in this article.
Note that the tissue_positions.csv contains the same information about the barcode whitelist and their slide coordinates as that contained in the barcodes folder that comes bundled with Space Ranger (spaceranger-x.y.z/lib/python/cellranger/barcodes where x.y.z represent the version number).
spatial_enrichment.csv: File in CSV format that contains table of Moran's I values for each feature when specific conditions are met:
The Moran's I value can be anywhere between -1 (perfectly dispersed) to 1 (perfectly enriched) but in biological samples values significantly below 0 are unexpected. A p-value is provided, as well as an adjusted p-value which is corrected using the Benjamini-Hochberg method for multiple comparisons.
From Space Ranger 2.0 onwards, the spatial_enrichment.csv file includes a Feature Type column the value for which is Gene Expression for single-library analysis.
# Gene Expression output $ cd /home/jdoe/runs/sampele345/outs/spatial $ column -s, -t < spatial_enrichment.csv | less -#2 -S Feature ID Feature Name Feature Type I P value Adjusted p value Feature Counts in Spots Under Tissue Median Normalized Average Counts Barcodes Detected per Feature ENSMUSG00000053310 Nrgn Gene Expression 0.849230495400066 0.0 0.0 140702 42.22927672729207 2645 ENSMUSG00000070570 Slc17a7 Gene Expression 0.8228828454463434 0.0 0.0 49919 15.927820652112793 2435 ENSMUSG00000046447 Camk2n1 Gene Expression 0.8112121614961633 0.0 0.0 126518 44.058800706638436 2698 ENSMUSG00000021948 Prkcd Gene Expression 0.8093848987347875 0.0 0.0 11675 4.552770117948635 1430 ENSMUSG00000061808 Ttr Gene Expression 0.8088005899881938 0.0 0.0 348686 76.73215387483816 2674 ENSMUSG00000041607 Mbp Gene Expression 0.7965984861055619 0.0 0.0 258769 112.84701279639039 2700 ENSMUSG00000035383 Pmch Gene Expression 0.7847484395395488 0.0 0.0 67814 17.005396313242215 1718 ENSMUSG00000024985 Tcf7l2 Gene Expression 0.7769492292430982 0.0 0.0 6546 2.564212465278306 1282 ENSMUSG00000032532 Cck Gene Expression 0.7605729556575779 0.0 0.0 67935 21.68537496001758 2585
barcode_fluorescence_intensity.csv: File in CSV format that is generated for any input image specified by --darkimage. This file contains the statistics about the fluorescence intensity which is calculated for every spot and every fluorescence channel in the input image.
If the channels were given custom names in the Loupe manual alignment session that produced the alignment JSON file, then the CSV header will use the given channel names. By default, channels are named channel1, channel2, etc.This file can be used for secondary analysis in R/Python and can also be combined with the tissue_positions.csv file.
$ cd /home/jdoe/runs/sampele345/outs/spatial $ column -s, -t < barcode_fluorescence_intensity.csv | less -#2 -S barcode in_tissue channel1_mean channel1_stdev channel2_mean channel2_stdev CGAGGATATTCAGAGC-1 0 4254.328631875882 59.72596599289198 2370.7574047954868 23.587614594004506 TCTGGTACTAATGCGG-1 0 4242.12976022567 61.31391702047626 2348.9830747531737 22.960436841214605 AGGTGACCATGACGCT-1 0 4237.420310296192 62.954555749425516 2330.277856135402 18.66240280182469 CAGAGGTGTACGTAGC-1 0 4219.421720733428 60.38810661890725 2323.2045133991537 20.876943016829802 TAGCTAGAAGATACAT-1 0 4202.056417489422 62.2343918192054 2309.320169252468 18.883665883691563 AGCACTGACCGGCTTA-1 0 4185.466854724965 61.848950071485 2300.589562764457 17.98861741514344 ATAACCTAAGGCGTCA-1 0 4149.37658674189 58.4482514520289 2292.1255289139635 19.824612785901802 TAGAACCTCAGTTAGA-1 0 4156.802538787024 64.86614573399514 2298.7390691114247 19.838300058320733 CCGACATAAGTATCAA-1 0 4194.483779971792 62.19636612986138 2313.277856135402 20.56714424032058 GACTACCAACTGGTCC-1 0 4204.746121297602 62.91221526144468 2328.5232722143865 54.82377031240396