Space Ranger2.0, printed on 04/14/2025
Space Ranger's pipelines analyze sequencing data produced from Visium Spatial Gene Expression for fresh frozen (FF) and formalin fixed paraffin embedded FFPE tissue samples. The workflow diagram shows all the required inputs and the corresponding spaceranger flags for FF tissue samples:
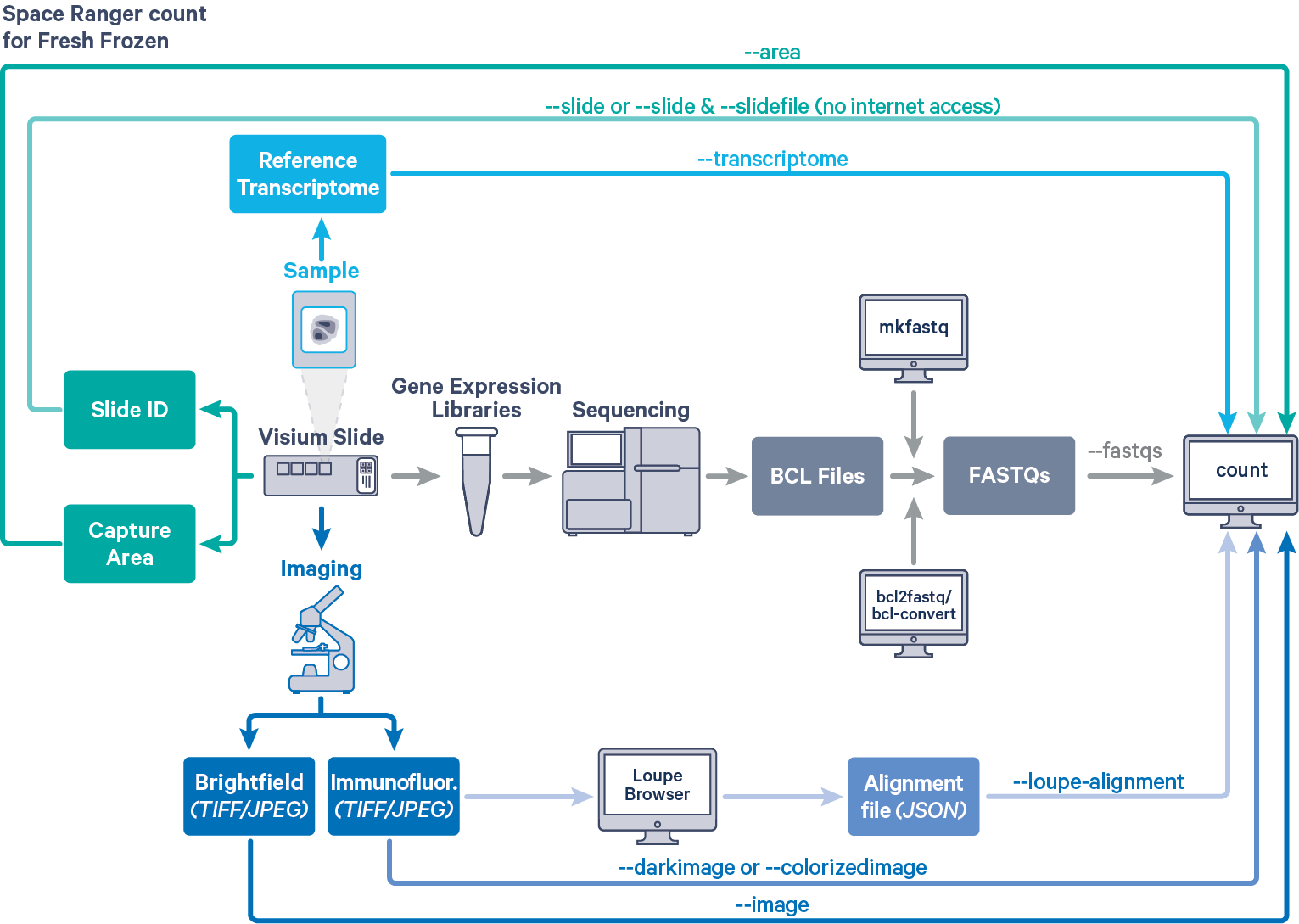
The spaceranger count is run on each individual Capture Area in the Visium slide. The required inputs are
--fastqs)
TIFF, QPTIFF or JPEG format
--image for brightfield image--darkimage for dark background fluorescence image--colorizedimage for composite colored fluorescence image--slide & --area if spaceranger has access to internet--slidefile, --slide & --area if spaceranger has no access to internet. The slide layout file is directly downloaded--unknown-slide if visium slide details are unknown--transcriptome)For a list of accepted arguments, see the Command Line Argument Reference below, or run spaceranger count --help.
Select the tab corresponding to the imaging workflow.
To generate spatial feature counts for a single library from fresh frozen (FF) sample using automatic fiducial alignment and tissue detection on a brightfield image input, run spaceranger count with the following arguments. The code in red indicates user specific inputs.
$ cd /home/jdoe/runs $ spaceranger count --id=sample345 \ #Output directory --transcriptome=/home/jdoe/refdata/GRCh38-2020-A \ #Path to Reference --fastqs=/home/jdoe/runs/HAWT7ADXX/outs/fastq_path \ #Path to FASTQs --sample=mysample \ #Sample name from FASTQ filename --image=/home/jdoe/runs/images/sample345.tiff \ #Path to brightfield image --slide=V19J01-123 \ #Slide ID --area=A1 \ #Capture area --localcores=8 \ #Allowed cores in localmode --localmem=64 #Allowed memory (GB) in localmode
To generate spatial feature counts for a single library fresh frozen (FF) sample using a fiducial alignment and tissue assignment
JSON file generated in Loupe Browser on a brightfield image input, run spaceranger count with the following arguments. The code in red indicates user specific inputs.
$ cd /home/jdoe/runs $ spaceranger count --id=sample345 \ #Output directory --transcriptome=/home/jdoe/refdata/GRCh38-2020-A \ #Path to Reference --fastqs=/home/jdoe/runs/HAWT7ADXX/outs/fastq_path \ #Path to FASTQs --sample=mysample \ #Sample name from FASTQ filename --image=/home/jdoe/runs/images/sample345.tiff \ #Path to brightfield image --slide=V19J01-123 \ #Slide ID --area=A1 \ #Capture area --loupe-alignment=sample345.json \ #Manual alignment file --localcores=8 \ #Allowed cores in localmode --localmem=64 #Allowed memory (GB) in localmode
The arguments to specify the fluorescence image can be either --darkimage or --colorizedimage depending on the image format. Details about the different supported fluorescence image formats are described in Image Recommendations.
Running preflight checks (please wait)... Checking sample info... Checking FASTQ folder... Checking reference... Checking reference_path Checking optional arguments... ...
A successful spaceranger count run concludes with a message similar to this:
Outputs:
- Run summary HTML: /home/jdoe/runs/sample345/outs/web_summary.html
- Outputs of spatial pipeline:
aligned_fiducials: /home/jdoe/runs/sample345/outs/spatial/aligned_fiducials.jpg
detected_tissue_image: /home/jdoe/runs/sample345/outs/spatial/detected_tissue_image.jpg
scalefactors_json: /home/jdoe/runs/sample345/outs/spatial/scalefactors_json.json
tissue_hires_image: /home/jdoe/runs/sample345/outs/spatial/tissue_hires_image.png
tissue_lowres_image: /home/jdoe/runs/sample345/outs/spatial/tissue_lowres_image.png
cytassist_image: null
aligned_tissue_image: null
tissue_positions: /home/jdoe/runs/sample345/outs/spatial/tissue_positions.csv
spatial_enrichment: /home/jdoe/runs/sample345/outs/spatial/spatial_enrichment.csv
barcode_fluorescence_intensity: null
- Run summary CSV: /home/jdoe/runs/sample345/outs/metrics_summary.csv
- Correlation values between isotypes and Antibody features: null
- BAM: /home/jdoe/runs/sample345/outs/possorted_genome_bam.bam
- BAM BAI index: /home/jdoe/runs/sample345/outs/possorted_genome_bam.bam.bai
- BAM CSI index: null
- Filtered feature-barcode matrices MEX: /home/jdoe/runs/sample345/outs/filtered_feature_bc_matrix
- Filtered feature-barcode matrices HDF5: /home/jdoe/runs/sample345/outs/filtered_feature_bc_matrix.h5
- Unfiltered feature-barcode matrices MEX: /home/jdoe/runs/sample345/outs/raw_feature_bc_matrix
- Unfiltered feature-barcode matrices HDF5: /home/jdoe/runs/sample345/outs/raw_feature_bc_matrix.h5
- Secondary analysis output CSV: /home/jdoe/runs/sample345/outs/analysis
- Per-molecule read information: /home/jdoe/runs/sample345/outs/molecule_info.h5
- Loupe Browser file: /home/jdoe/runs/sample345/outs/cloupe.cloupe
- Feature Reference: null
- Target Panel file: null
- Probe Set file: null
Pipestance completed successfully!
The output of the pipeline is contained in a folder named with the sample ID specified using --id (e.g. sample345). If this folder
already exists, in a rerun with the original parameters, spaceranger will assume it is an existing pipestance and attempt to resume running it. The subfolder named outs contains the main pipeline output files.
Once spaceranger count has successfully completed, you can further explore the results by
.cloupe file in
Loupe Browser| Argument | Description |
|---|---|
--id | Required. A unique run ID string (e.g., sample345). The name is arbitrary and will be used to name the directory containing all pipeline-generated files and outputs. Only letters, numbers, underscores, and hyphens are allowed (maximum of 64 characters). |
--fastqs | Required. Either: Path of the fastq_path folder generated by spaceranger mkfastq (such as /home/jdoe/runs/HAWT7ADXX/outs/fastq_path). This contains a directory hierarchy that spaceranger count will automatically traverse.- OR - Any folder containing fastq files, for example if the fastq files were generated by a service provider and delivered outside the context of the mkfastq output directory structure. Can take multiple comma-separated paths, which is helpful if the same library was sequenced on multiple flowcells. Doing this will treat all reads from the library, across flowcells, as one sample. |
--sample | Optional. Sample name as specified in the sample sheet supplied to spaceranger mkfastq.
Can take multiple comma-separated values, which is helpful if the same library was sequenced on multiple flowcells and the sample name used (and therefore fastq file prefix) is not identical between them. Doing this will treat all reads from the library, across flowcells, as one sample. Allowable characters in sample names are letters, numbers, hyphens, and underscores. |
--transcriptome | Required. Path to the Space Ranger compatible transcriptome reference. (such as /home/jdoe/GRCh38-2020-A) |
--image | Required for brightfield image input. Brightfield tissue H&E image in JPEG or TIFF format. For more information on image file constraints, encoding and formats refer the Image Recommendations. |
--darkimage | Required for dark background fluorescence image input. Multi-channel, dark-background fluorescence image as either a single, multi-layer TIFF file or as multiple TIFF or JPEG files (which can provided as a comma separated list or by invoking the --darkimage parameter multiple times). Details on image file constraints, encoding and formats are described in the Image Recommendations section. |
--colorizedimage | Required for color composite fluorescence image input. A color composite of one or more fluorescence image channels saved as a single-page, single-file color TIFF or JPEG. Please see the Image Recommendations section for information on input image file formats. |
--slide | Visium slide serial number. Refer to the Slide Parameters for information on supported slide versions. Required unless --unknown-slide is passed. |
--area | Visium capture area identifier. Required unless --unknown-slide is passed. Options for Visium are A1, B1, C1, D1. |
--slidefile | Slide layout file indicating capture spot and fiducial spot positions. Required when no internet access unless --unknown-slide is passed. |
--unknown-slide | Optional. Set this if the slide serial number and area identifier are unknown. The value passed to this depends of the slide version. Possible value for single-library FF workflow is visium-1. Setting this will cause Space Ranger to use default spot positions. Not compatible with --slide, --area, or --slidefile. Refer to the Slide Parameters for more information. |
--reorient-images | Default:true. Whether the pipeline should rotate and mirror the image to align the upright hourglass fiducial pattern in the top left corner. Set --reorient-images=false if you are certain your image is already oriented with the upright hourglass in the top left corner. |
--loupe-alignment | Required for manual alignment. Alignment file produced by the manual Loupe alignment step. A --image, --darkimage or --colorizedimage, the same image(s) that were used for manual alignment, must be supplied in this case. |
--no-bam | Optional. Add this flag to skip generating BAM file. This will reduce the total computation time for the pipestance and the size of the output directory. If unsure, we recommend not using this option, as BAM files can be useful for troubleshooting and downstream analysis. |
--nosecondary | Optional. Disable secondary analysis, such as dimensionality reduction, clustering, spatial enrichment calculation (Moran's I) and visualization. |
--r1-length | Optional. Limit the length of the input R1 sequence to the first N bases where N is the value supplied by the user. Note that the length includes the Barcode and UMI sequences so do not set this below 28. This and --r2-length are useful for determining the optimal read length for sequencing. |
--r2-length | Optional. Limit the length of the input R2 sequence to the first N bases where N is the value supplied by the user. |
--lanes | Optional. Lanes associated with this sample. |
--localcores | Recommended when run in localmode. Restricts spaceranger to use specified number of cores to execute pipeline stages. By default, spaceranger will use all of the cores available on your system. |
--localmem | Recommended when run in localmode. Restricts spaceranger to use specified amount of memory (in GB) to execute pipeline stages. By default, spaceranger will use 90% of the memory available on your system. |