Cell Ranger5.0, printed on 04/02/2025
Loupe V(D)J Browser v4.0 and later allows you to load .vloupe files from the cellranger aggr pipeline for multi-sample comparisons.
For a demo of the different comparisons available, download the two-sample PBMC .vloupe file.
Upon opening the aggregated file in Loupe V(D)J Browser, the views are the same as the ones produced for a single sample file, however, every view can contain data from multiple samples.
Notice that the Filters pop-up window now has a 'Sample' filter.
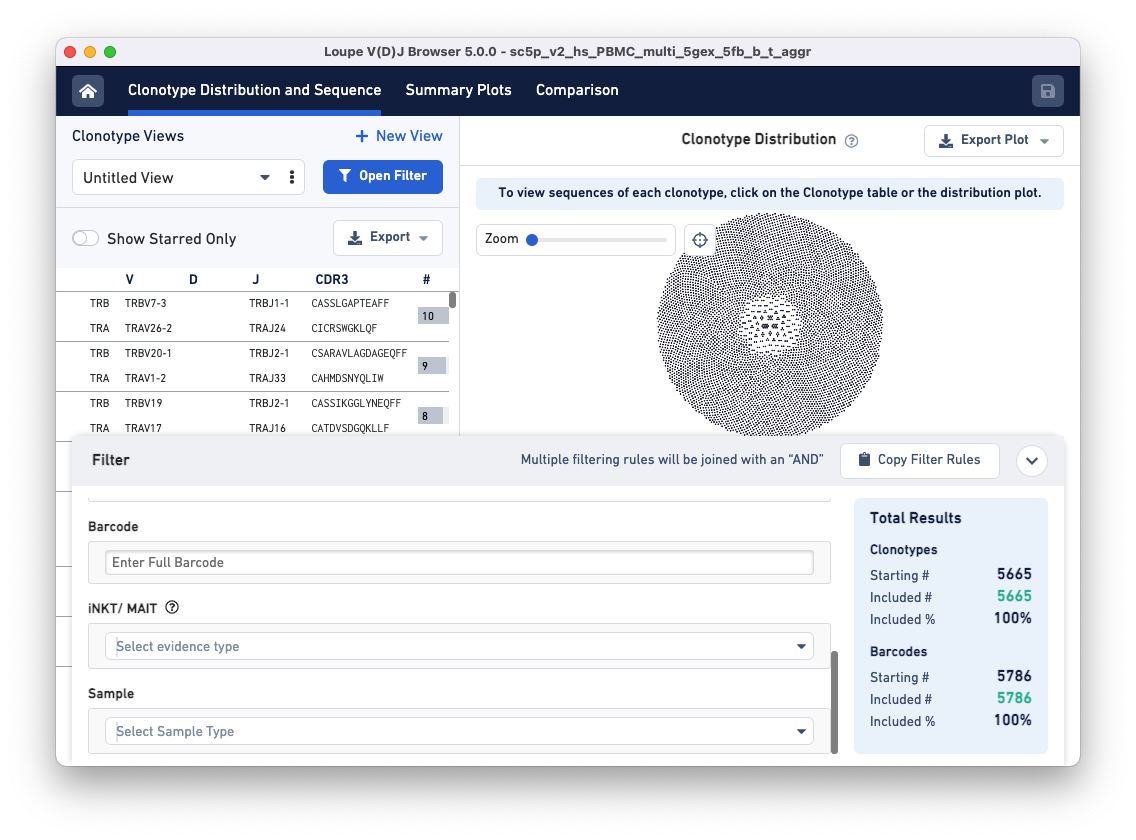
Clicking on the Select Sample Type drop-down brings up a list of options for dividing the barcodes in the file into samples. These sample name options are based on a column in the aggregation CSV, an input file for the cellranger aggr pipeline. The default options are Donor, Origin, and Library ID, but custom fields like 'Time Points' or 'Disease Status' may also be provided.
For details on the definitions of the default groups look at the Cell Ranger glossary.
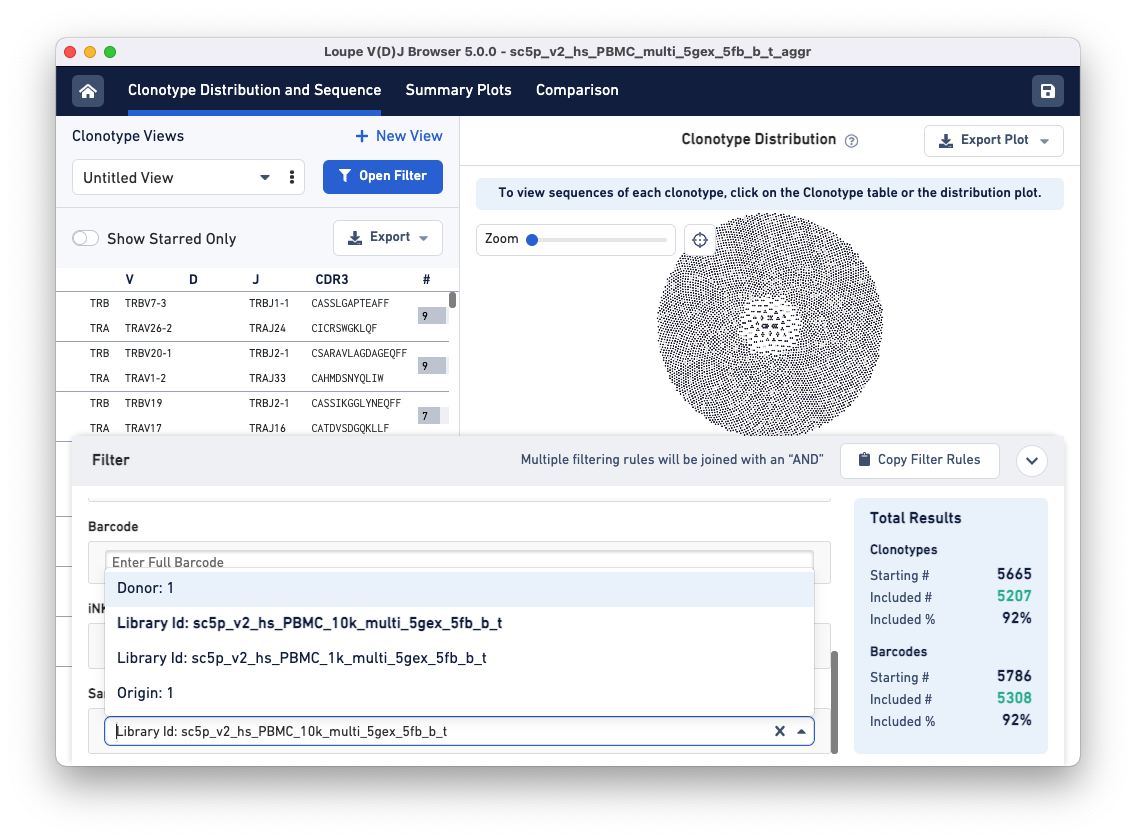
To compare the samples in detail, click on the Comparison tab.
The default view is the Sample Table per donor in the aggregated dataset. The table lists all the loaded V(D)J samples with their Barcode and Clonotype counts. There is also a sparkline of each sample's top 100 clonotype abundances. The table and other charts accessible from the drop-down menu show samples divided by sample groups (donor, origin, library ID, or custom groups).
In this example, there is only one row because the two aggregated libraries come from the same donor:
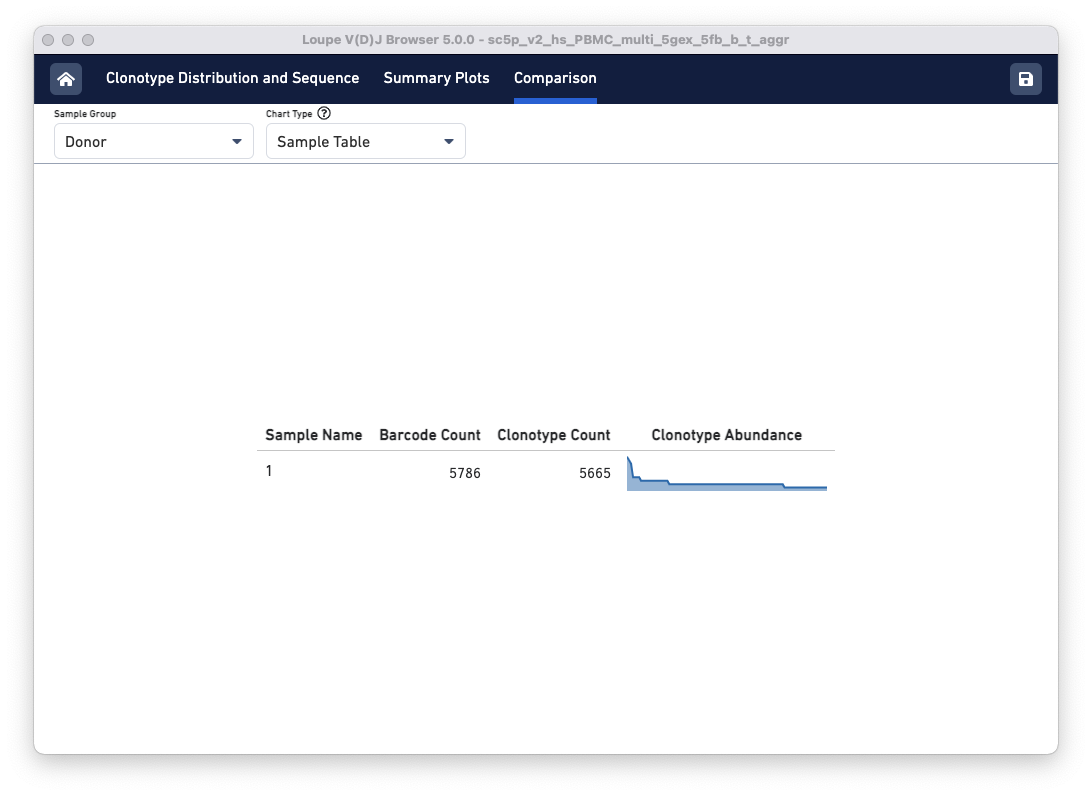
Above the sample table are all other comparison charts is the control bar. The Sample Group selector can be used to divide the data views in different ways. The tutorial file has the same donor and origin but comes from two different libraries, so toggle to the Library Id option. Using the Chart Type selector, select Top Clonotype Overlap to see the top 10 clonotypes in one library compared to the same clonotypes in the second library. You can change the library whose top 10 Clonotypes you wish to compare using the Top 10 Clonotypes from drop-down menu on the top right.
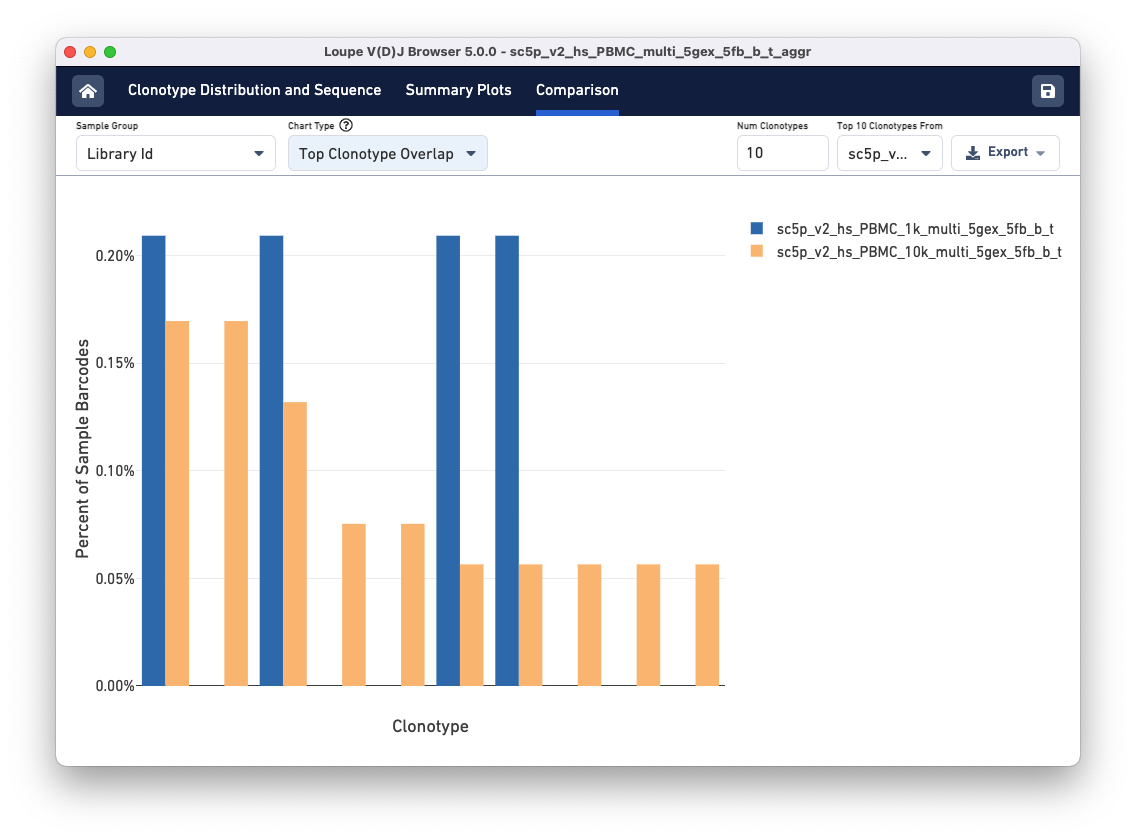
The top 10 clonotypes from the 10k sample (orange) are also fairly abundant in the 1k sample (blue). You can change the number of clonotypes compared by editing the Num Clonotypes value in the control bar.
The Clonotype Frequency chart gives you a feel for how closely the clonotypes of two samples map onto each other. The x-axis represents the frequency of a particular clonotype in sample A, and the y-axis represents the frequency of a particular clonotype in sample B. The size of the circles at each point represents the number of clonotypes that have x barcodes in sample A and y barcodes in sample B.
To get a feel for this, compare the two samples by library ID. The mutual frequencies are lined up vertically in the center of the chart. In this case, the green sample primarily has clonotypes with one barcode, which is where that line falls, but no overlap in clonotypes with multiple barcodes.
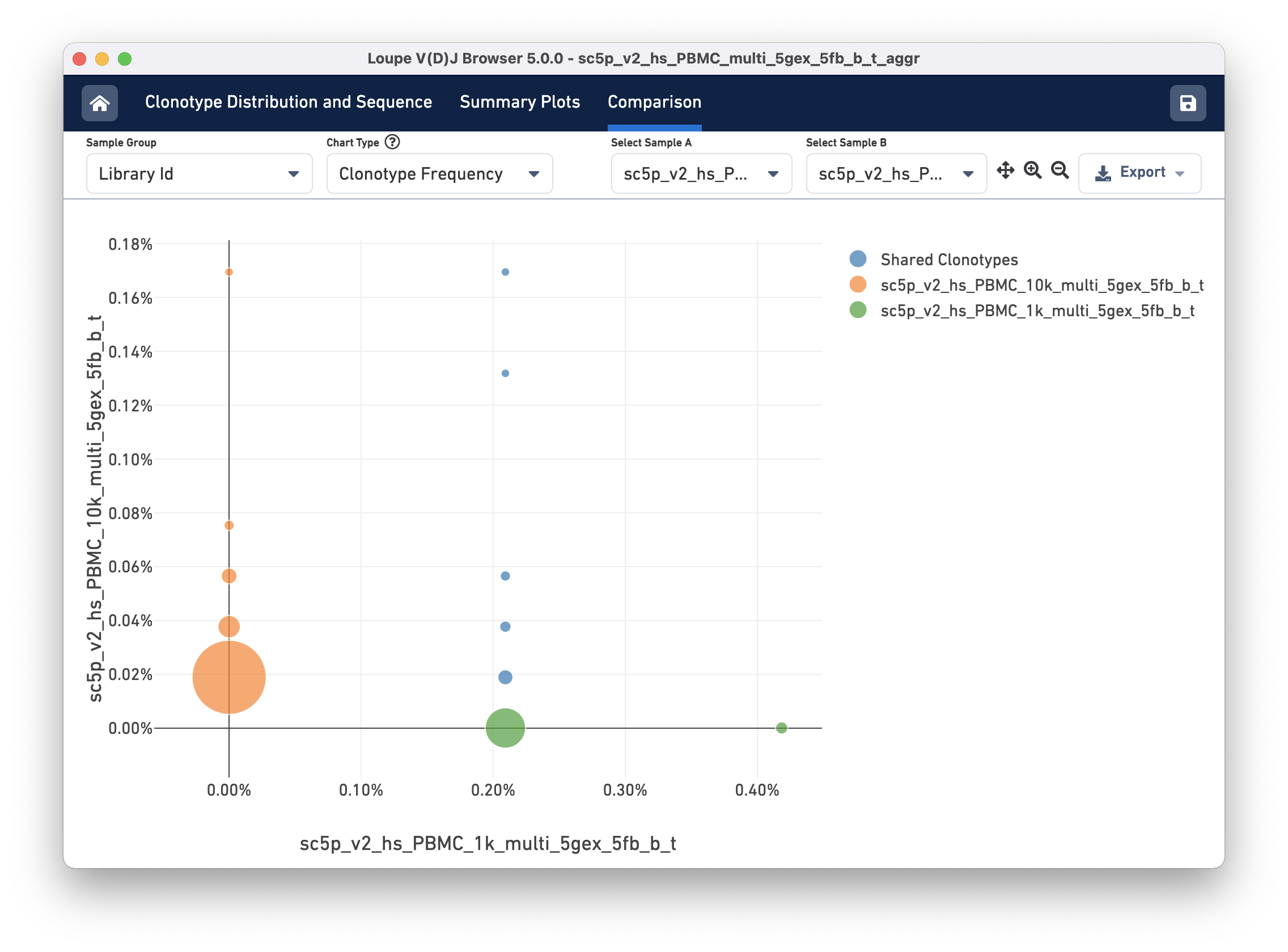
You can use the zoom in/out and autoscale buttons on the control bar to zoom into regions of the chart in more detail, and drag the chart around with your mouse.
Finally, you can compare the relative frequencies of V, D, J, and C genes across multiple samples by selecting one of those four charts in the chart type selector. For each gene usage chart, you can either look at all chain types or limit to a single chain type through the Chain Type selector.
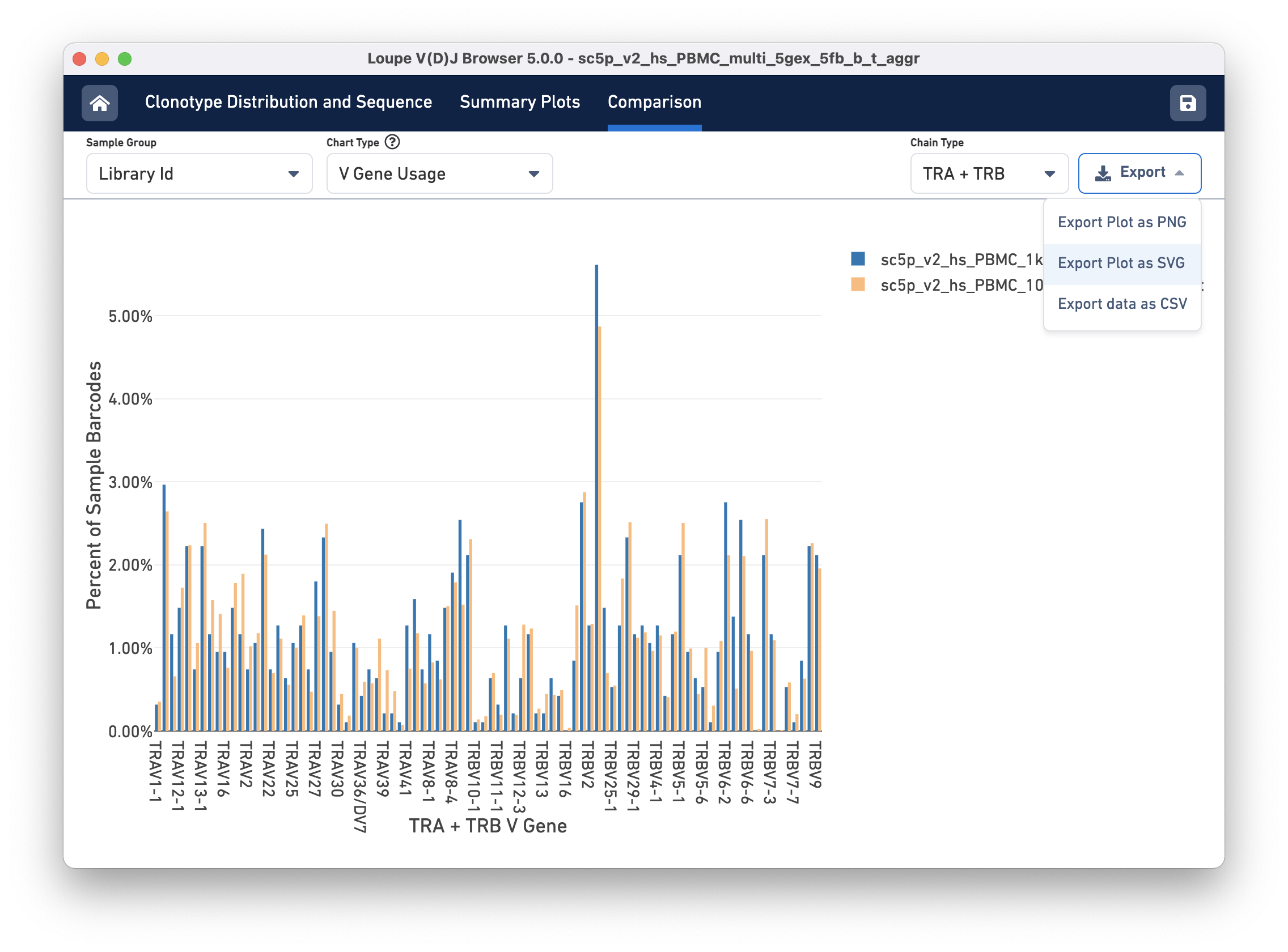
Most charts have an export button, where the chart image can be exported to SVG or PNG, or the underlying data can be exported to a CSV file (shown in the image above). When the file has a large number of samples, another selector will appear that can be used to select a subset of samples to view throughout the charts.
This concludes the Loupe V(D)J Browser tutorial. We hope that you are comfortable enough now to use this tool on your Chromium Single Cell 5′ V(D)J data. If you have matching 5′ Gene Expression data, the Loupe V(D)J Browser and Loupe Browser are more powerful together. See the integrated V(D)J and gene expression analysis tutorial.
If you encounter difficulties or have general questions about the software, email support@10xgenomics.com with your questions.