Cell Ranger6.4, printed on 03/29/2025
|
Documentation and downloads for Loupe Browser 7.0 are now available on our redesigned software support site. This site is being maintained temporarily to support older versions of Cell Ranger and Loupe Browser, and all versions of Loupe V(D)J Browser. |
This set of tutorials review the major analysis capabilities Loupe Browser provides for analyzing 3' Single Cell Gene Expression, Feature Barcode data, and 5' V(D)J and Single Cell Gene Expression data.
To use Loupe Browser, follow the directions on the Downloads page to download and install the software on either macOS or Windows.
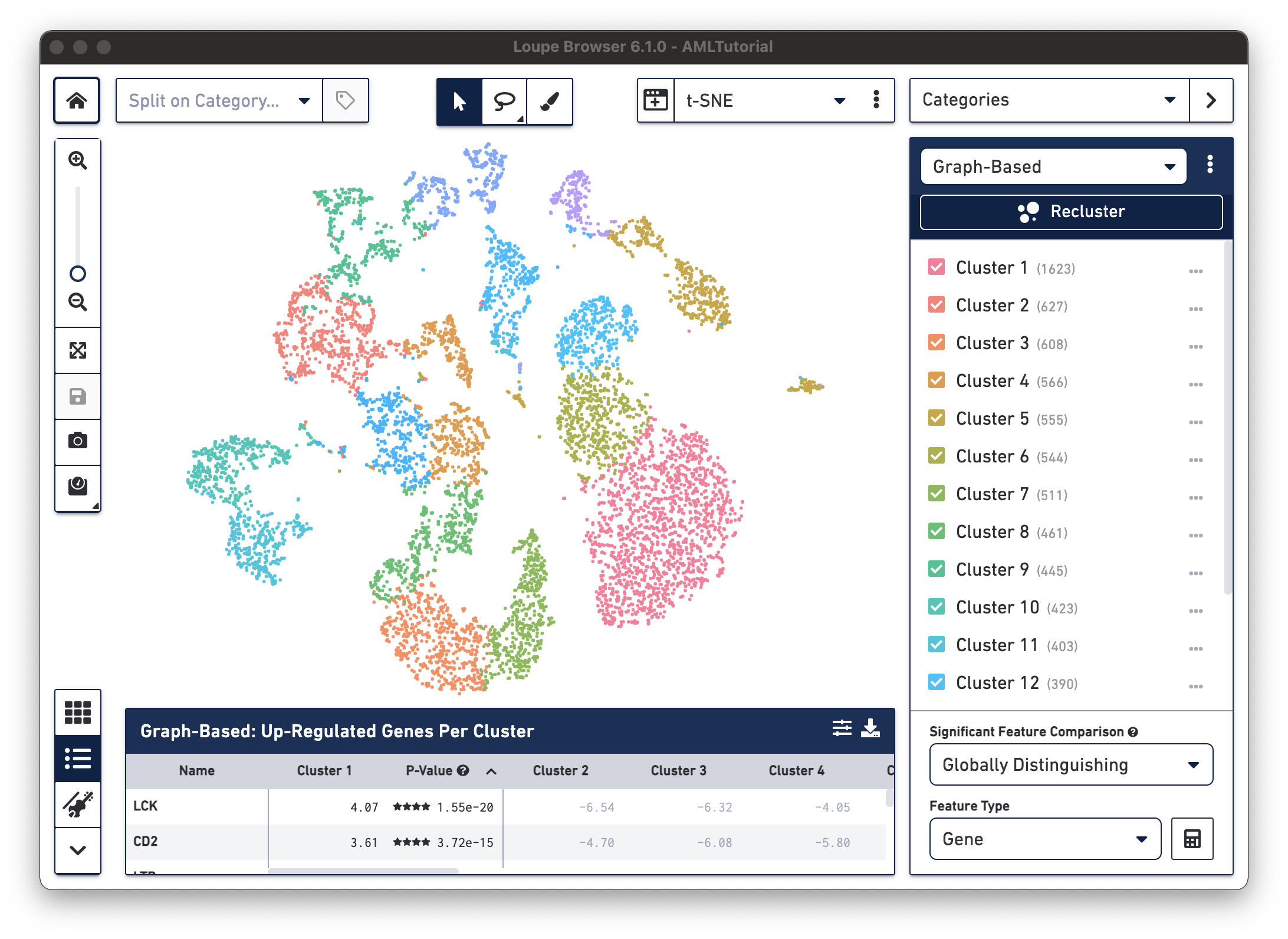
Open Loupe Browser by double-clicking on the application icon. Then click on the
AMLTutorial file from the list of Recent Files. You should see a screen with the t-SNE projection for the cells from
the AML dataset.
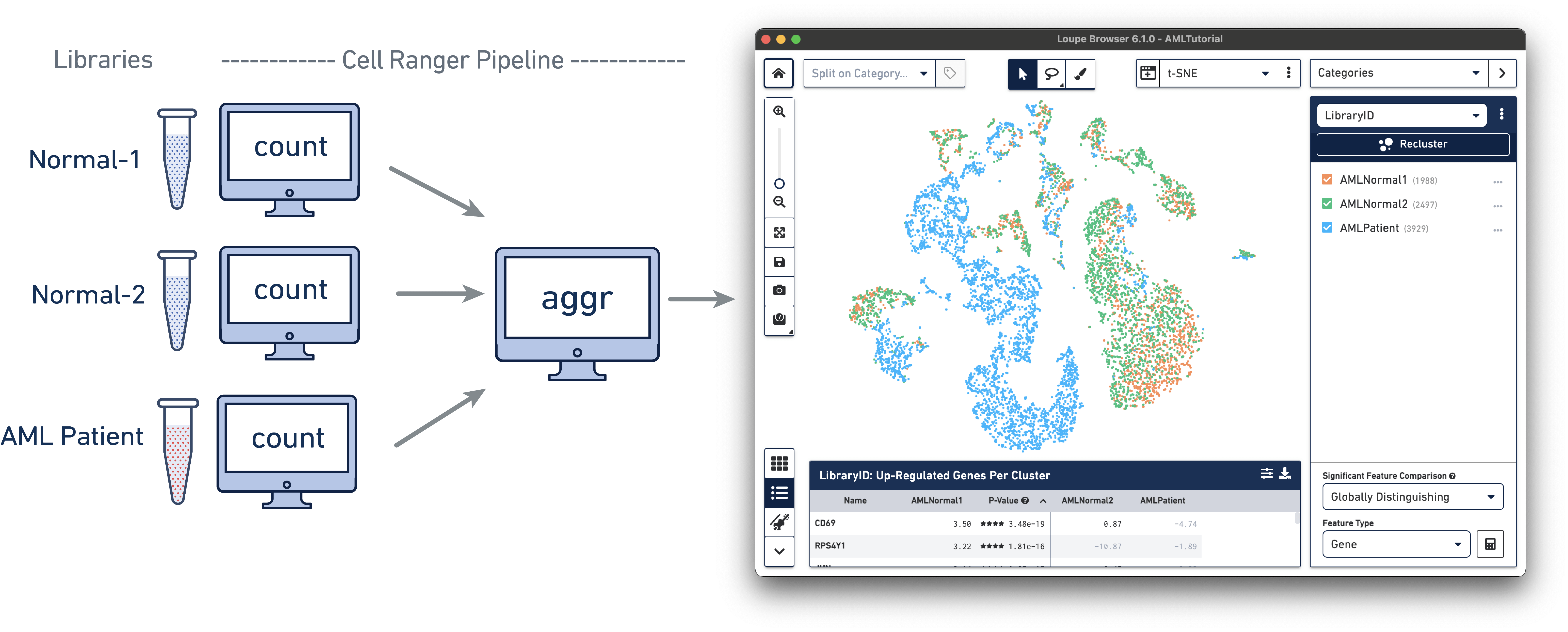
Once you have installed Loupe Browser, you can follow the tutorials using the AML example dataset:
The AML Tutorial dataset contains the results of a Cell Ranger aggr pipeline analysis run for three samples, including two healthy control samples of frozen human bone marrow mononuclear cells and a pre-transplant sample from a patient with acute myeloid leukemia (AML). This dataset was generated in collaboration with the Fred Hutchinson Cancer Research Center, and referenced in the Nature Communications publication, "Zheng et al, Massively parallel digital transcriptional profiling of single cells" (2017; doi:10.1038/ncomms14049).
Starting with Loupe Browser v6.1, users may choose to share anonymized non-biological usage data (i.e. no identifiable sample information such as files, cluster names, or any open text fields are tracked) to improve Loupe Browser performance. Sharing of usage data is optional, and users can change whether they want to share usage data any time. The user preference for usage data sharing will carry over within a major version (e.g. all 6.Y versions) but the user will be asked to confirm their preferences for each major version update.
In case you wish to change the usage data sharing setting, click Manage Usage Data from the Help menu and change the preference in the pop-up window.