Cell Ranger7.1, printed on 03/29/2025
Job submission mode is one of four primary ways of running Cell Ranger. To learn about the other approaches, please go to the computing options page.
Cell Ranger can be run in job submission mode, by treating a single node from the cluster like a local server. This leverages existing institutional hardware for pipeline analysis.
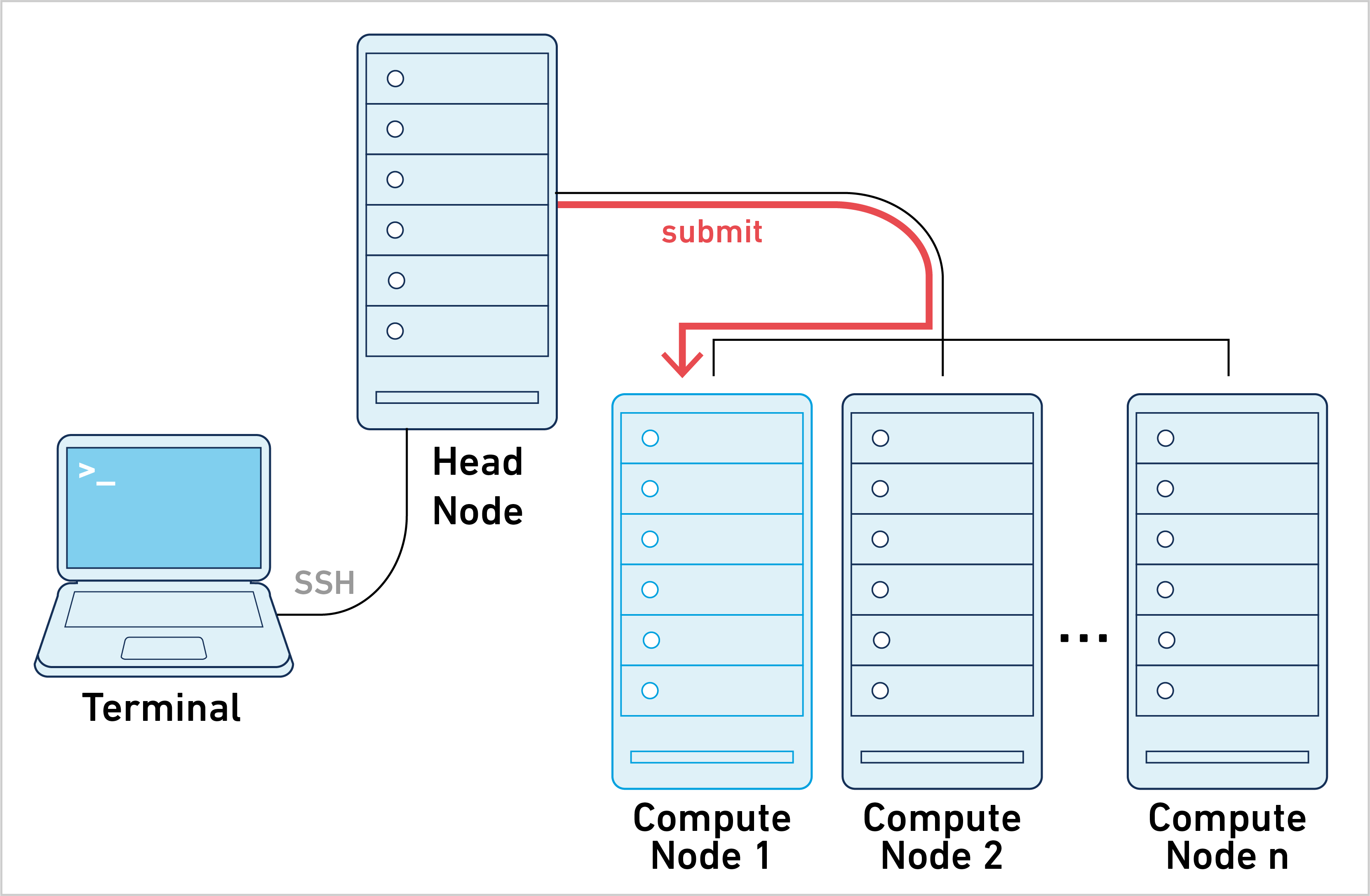
Running cellranger commands in this mode is typically achieved by submitting a job script to the cluster. The process is as follows:
Step 1. Select a job scheduler tab below to see a sample job script.
| 10X does not officially support Slurm or Torque/PBS. However, many customers have successfully used Cell Ranger with those job schedulers in job submission mode. Sample job scripts are shown below. |
Step 2. Copy the sample job script to a text editor, and replace all the variables enclosed within braces (e.g. {JOB_NAME} or {STDOUT_LOG}) with literal names, paths, and values. The last line, {PIPELINE_CMD} , is replaced by a Cell Ranger command, specifying --jobmode=local, --localcores={NUM_THREADS} and --localmem=0.9*{MEM_GB} to restrict CPU and memory use that's consistent with the directives in the job script. For example, if the job script requests 8 cores and 64GB of memory from the cluster, then the Cell Ranger command should include --jobmode=local --localcores=8 --localmem=57.
| Since cluster setup and policy vary greatly from one institution to another, the sample job scripts provided below should be treated as a starting point. Consult with your local cluster administrator to see if additional directives are required. |
#!/usr/bin/env bash
#
# =============================================================================
# Setup Instructions
# =============================================================================
#
# 1. Substitute {PE_NAME} below with name of your cluster's shared-memory
# parallel environment. If your cluster does not have a parallel environment,
# delete this line. However, the job will run with only 1 thread.
#
# =============================================================================
# Job Script
# =============================================================================
#
#$ -N {JOB_NAME}
#$ -V
#$ -pe {PE_NAME} {NUM_THREADS}
#$ -cwd
#$ -l mem_free={MEM_GB}G
#$ -o {STDOUT_LOG}
#$ -e {STDERR_LOG}
#$ -S "/usr/bin/env bash"
{PIPELINE_CMD}
Step 3. If this job script is saved as runJob.bash, then the following command submits the job to the cluster:
$ qsub runJob.bash
#!/usr/bin/env bash
#
# =============================================================================
# Job Script
# =============================================================================
#
#BSUB -J {JOB_NAME}
#BSUB -n {NUM_THREADS}
#BSUB -o {STDOUT_LOG}
#BSUB -e {STDERR_LOG}
#BSUB -R "rusage[mem={MEM_MB}]"
#BSUB -R span[hosts=1]
{PIPELINE_CMD}
Step 3. If this job script is saved as runJob.bash, then the following command submits the job to the cluster:
$ bsub runJob.bash
#!/usr/bin/env bash
#
#!!! This is not officially supported by 10x
#
# =============================================================================
# Job Script
# =============================================================================
#
#SBATCH -J {JOB_NAME}
#SBATCH --export=ALL
#SBATCH --nodes=1 --ntasks-per-node={NUM_THREADS}
#SBATCH --signal=2
#SBATCH --no-requeue
### Alternatively: --ntasks=1 --cpus-per-task={NUM_THREADS}
### Consult with your cluster administrators to find the combination that
### works best for single-node, multi-threaded applications on your system.
#SBATCH --mem={MEM_GB}G
#SBATCH -o {STDOUT_LOG}
#SBATCH -e {STDERR_LOG}
{PIPELINE_CMD}
Step 3. If this job script is saved as runJob.bash, then the following command submits the job to the cluster:
$ sbatch runJob.bash
#!/usr/bin/env bash
#
#!!! This is not officially supported by 10x
#
# =============================================================================
# Job Script
# =============================================================================
#
#PBS -N {JOB_NAME}
#PBS -V
#PBS -l nodes=1:ppn={NUM_THREADS}
#PBS -l mem={MEM_GB}gb
#PBS -o {STDOUT_LOG}
#PBS -e {STDERR_LOG}
cd {JOB_WORKDIR}
{PIPELINE_CMD}
Step 3. If this job script is saved as runJob.bash, then the following command submits the job to the cluster:
$ qsub runJob.bash