Cell Ranger3.1, printed on 03/31/2025
Loupe Cell Browser is a desktop application for Windows and MacOS that allows you to quickly and easily visualize and analyze 10x Chromium™ Single Cell 5′ and 3′ gene expression data. It is optimized for finding significant genes, identifying cell types, and exploring substructure within cell clusters. Loupe is named for a jeweler's loupe, which is used to inspect gems.
| 10x Chromium | Sequencer | 10x Cell Ranger | 10x Loupe Cell Browser | |
|---|---|---|---|---|
| Cell Suspension |
Barcoding & Library Construction |
Sequence library | Pipelines | Analysis & Visualization |
 |
 |
 |
 |
 |
Loupe Cell Browser opens .cloupe files generated by the Cell Ranger 1.3 (or later) pipelines, or by cellranger mkloupe for older pipelines. You may also import a sample's corresponding V(D)J clonotype data from Cell Ranger 2.1 or later. Once your data is in Loupe Cell Browser, you can rapidly explore and gain insights from the data without writing a line of code:
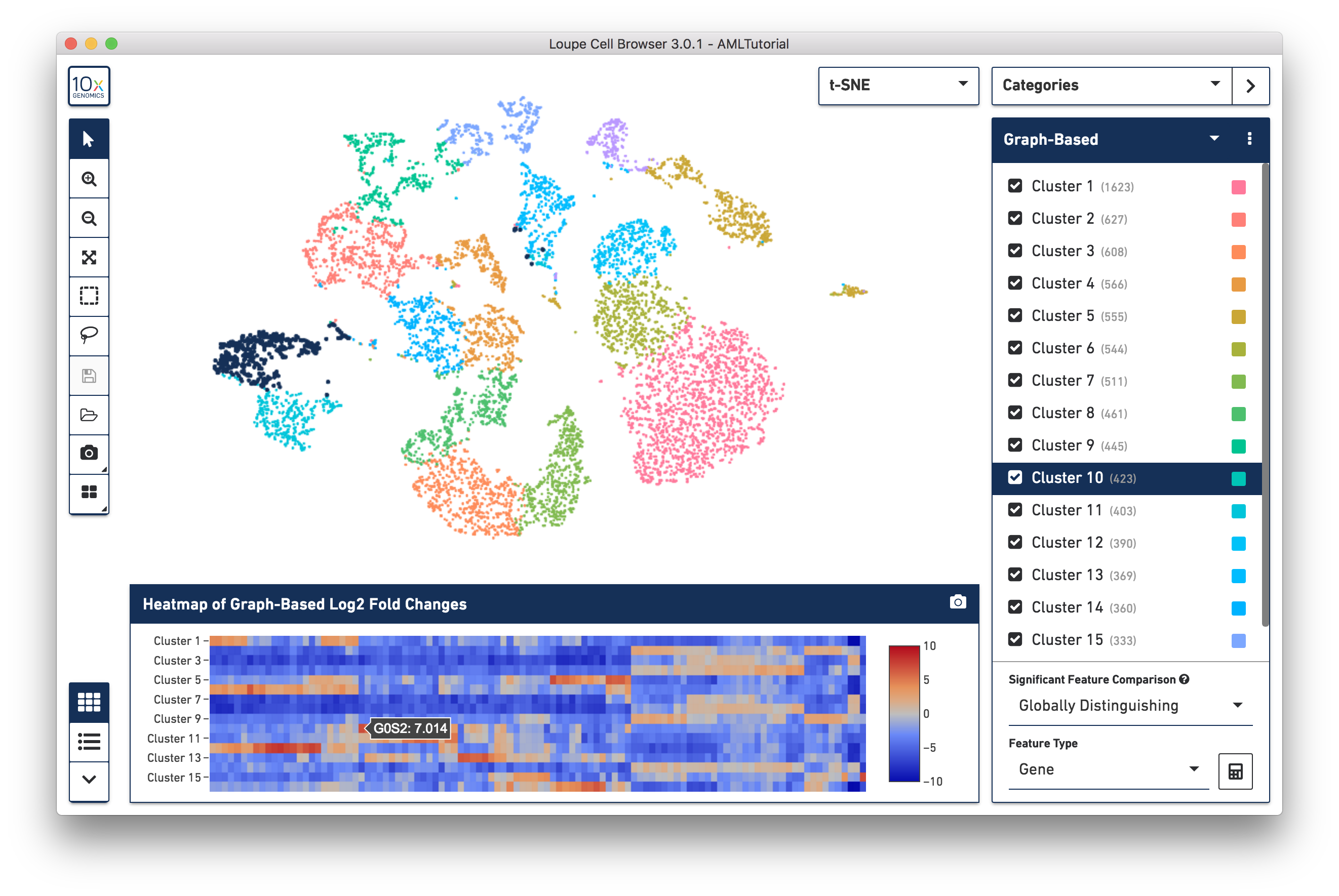
Loupe Cell Browser is built to accelerate the following applications:
Finding Significant Genes
Determine genes that uniquely characterize clusters at the press of a
button.
Identifying Cell Types
Use gene lists and the gene expression view to locate different cell types and
functional groups.
Exploring Substructure
Create custom clusters and use differential expression tools to identify even
smaller subgroups in your data.
Exploring Cell Subtypes
Use the filter panel to create complex boolean filters to find cell subtypes
in your dataset.
Integrated Gene Expression and V(D)J Analysis
Combine gene expression and immune repertoire data from the same sample to
gain new insights about complex disease.
Sharing Results
Save genes of interest, export data tables, and capture screenshots of your
single-cell data.
To walk through the features and uses of Loupe Cell Browser, start the tutorial.