Cell Ranger7.1, printed on 04/12/2025
The cellranger count pipeline outputs an interactive summary HTML file named web_summary.html that contains summary metrics and automated secondary analysis results. If an issue was detected during the pipeline run, an alert appears on this page. More information about alerts is provided in the Cell Ranger troubleshooting documentation. For more detail, please read the Interpreting Cell Ranger Web Summary Files for Single Cell Gene Expression Assays technical note.
The cellranger count web summary has Summary and Gene Expression tabs, and an additional Antibody tab for Antibody Capture analysis. Similar web summaries are also output from the cellranger reanalyze and cellranger aggr pipelines. aggr web summaries are described here.
The run summary from cellranger count can be viewed by clicking Summary in the top left tab of the HTML file. The summary metrics describe sequencing quality and various characteristics of the detected cells.
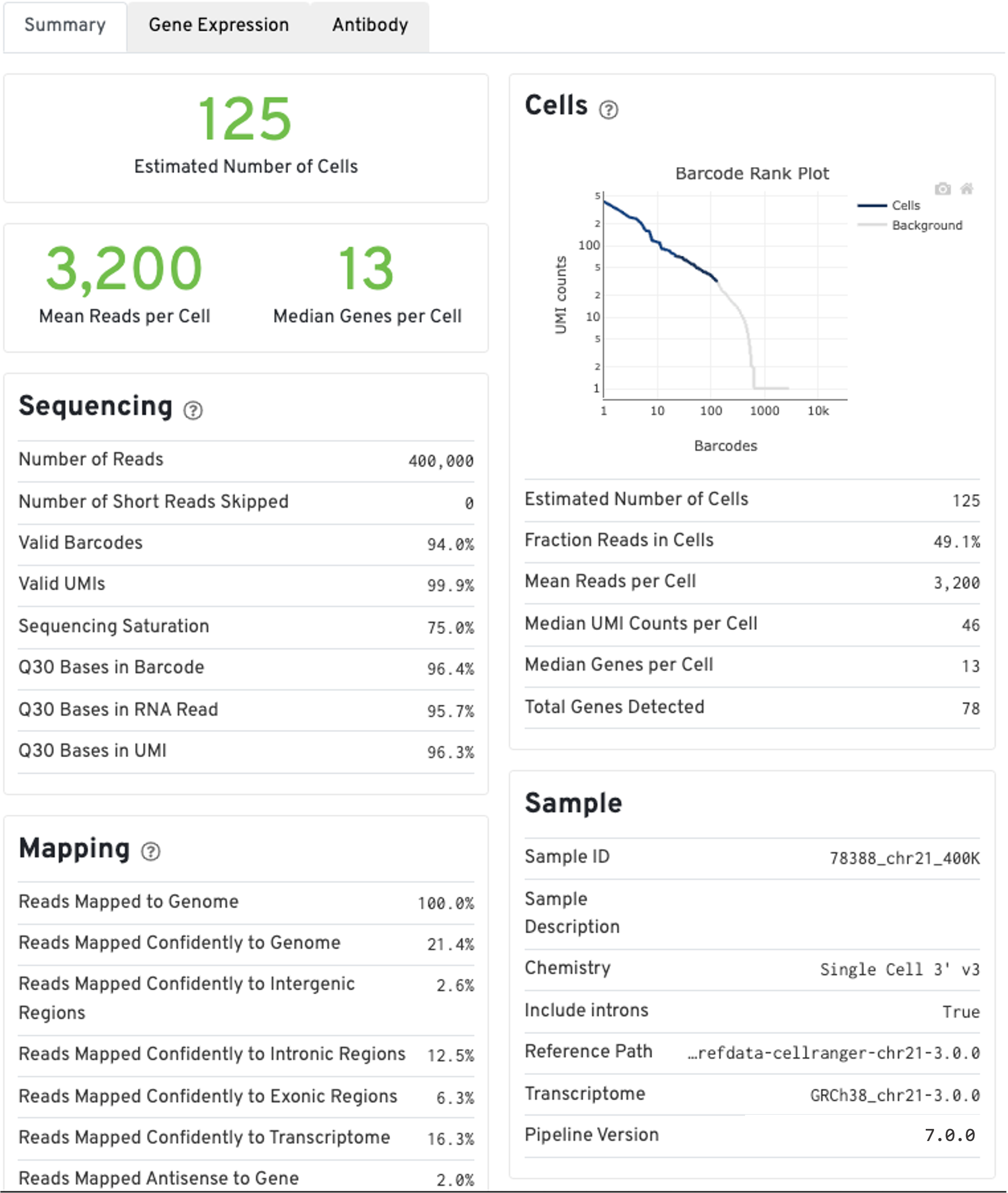
The estimated number of cells detected, mean reads per cell, and median genes detected per cell are prominently displayed near the top of the page.
Click the ? icons next to the Sequencing, Mapping, and Cells sections to display information about each metric in the dashboard.
The GEX Barcode Rank Plot under the Cells dashboard shows the distribution of barcode counts and which barcodes were inferred to be associated with cells. The y-axis is the number of UMI counts mapped to each barcode and the x-axis is the number of barcodes below that value. A steep drop-off is indicative of good separation between the cell-associated barcodes and the barcodes associated with empty partitions. Since barcodes can be associated with cells based on their UMI count or by their RNA profiles, some regions of the graph can contain both cell-associated and background-associated barcodes. The color of the graph represents the local density of barcodes that are cell-associated. See this Guided Tour of the Barcode Rank Plot for more details.
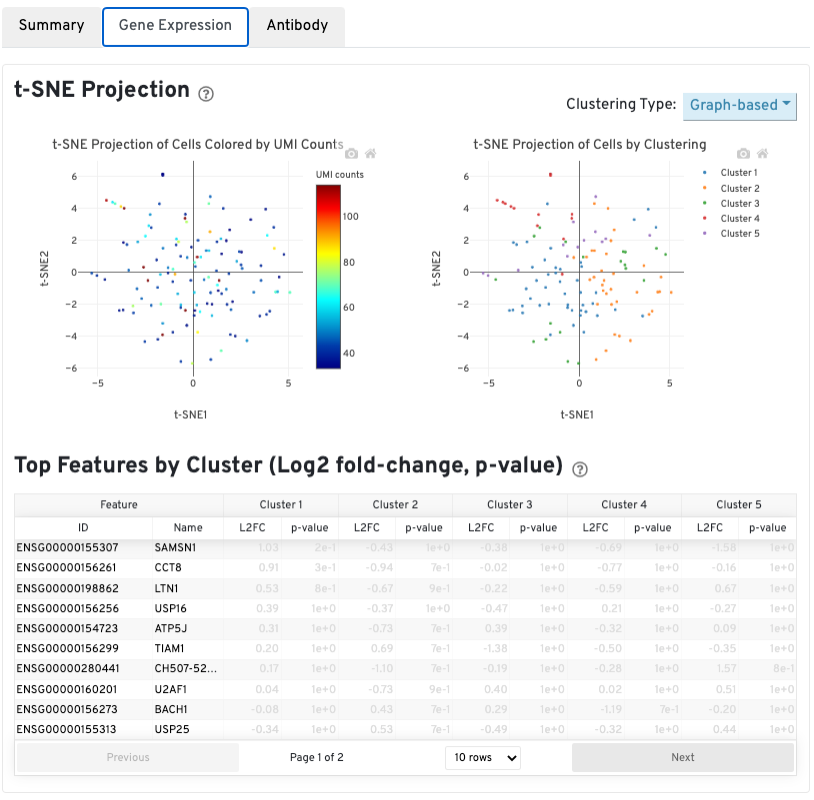
The automated secondary analysis results can be viewed by clicking the Gene Expression tab in the top left corner. Click the ? icons next to each section title to display information about the secondary analyses shown in the dashboard.
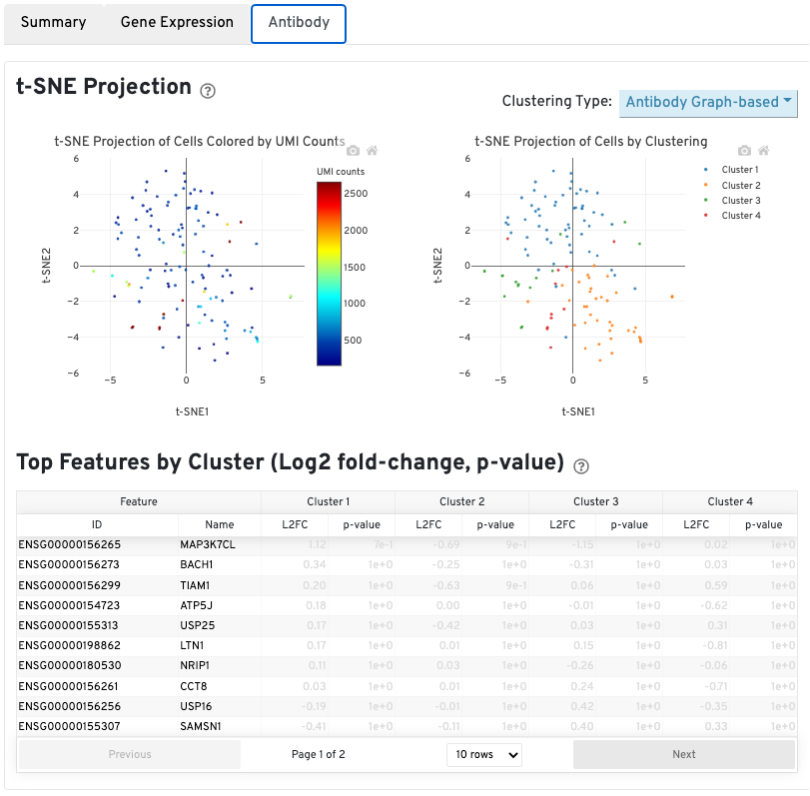
The t-SNE Projection section shows the data reduced to two dimensions, colored by UMI count (left) or clustering (right). It is a good starting point to explore structure in the data. The projection colored by UMI counts is indicative of the RNA content of the cells and often correlates with cell size - redder points are cells with more RNA in them. For the projection colored by clustering results, select the type of clustering analysis to display from the drop-down button on the upper right (Graph-based by default) - change the category to vary the type of clustering and/or number of clusters (K=2-10) that are assigned to the data.
The Top Features By Cluster table shows which genes are differentially expressed in each cluster relative to all other clusters (Graph-based by default). To find the genes associated with a particular cluster, click the L2FC or p-value column headers associated with a given cluster number to sort the table by a specific cluster.
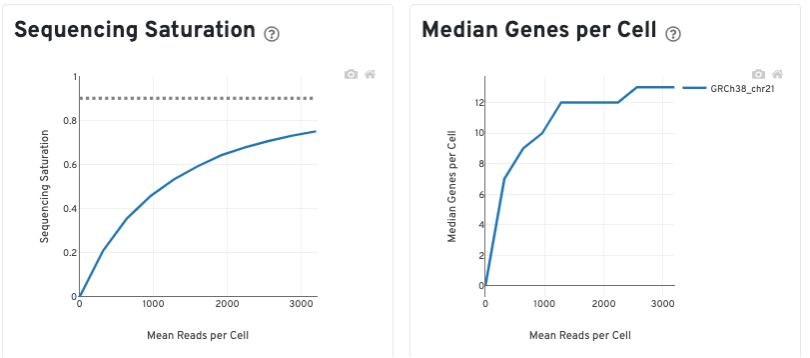
The Sequencing Saturation plot shows the effect of decreased sequencing depth on sequencing saturation, which is a measure of the fraction of library complexity that was observed. The right-most point on the line is the full sequencing depth obtained in this run.
Similarly, the Median Genes per Cell plot shows the effect of decreased sequencing depth on median genes per cell, which is a way of measuring data yield as a function of depth. The right-most point on the line is the full sequencing depth obtained in this run.
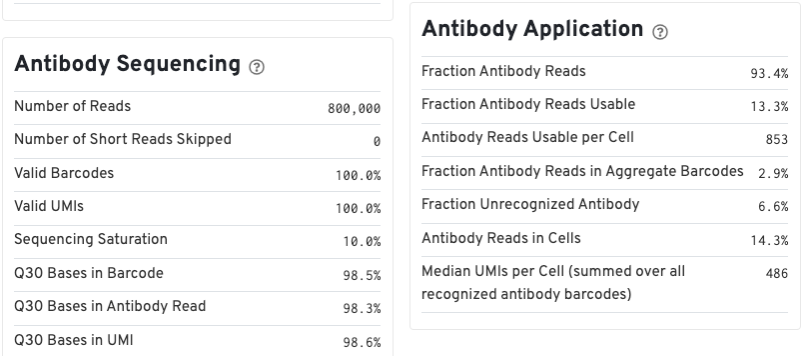
For Feature Barcode data, new metrics specific to Feature Barcode libraries will appear in the Summary tab in a new dashboard (e.g., Antibody Application, CRISPR Application), as well as on an Antibody tab for Antibody Capture analysis.
The Gene Expression tab view for Feature Barcode experiments contains the same secondary analysis as gene expression only experiments, and uses only the gene expression part of the feature-barcode matrix for dimensionality reduction, clustering, and differential gene expression analysis. Click the question mark ? to read full descriptions for each section.
Below is an example web summary for Gene Expression and Antibody Capture libraries. The Summary tab will have these additional dashboards:
The Antibody tab will have t-SNE projection plots, a Top Features table, Distribution of Antibody Counts plot, and a Histogram of Antibody counts:
The Distribution of Antibody Counts plot in Cell Ranger v7.1 and later shows the relative composition of antibody counts for antibodies with at least one UMI. The box size is proportional to the fraction of total UMIs from cell barcodes derived from that antibody. Hovering over a box provides more information about the metrics displayed. In this example, the first blue box shows that at least one CD45RA antibody UMI was detected in 535 cells:
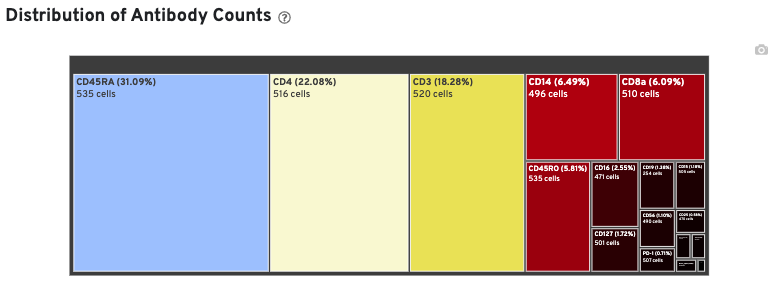

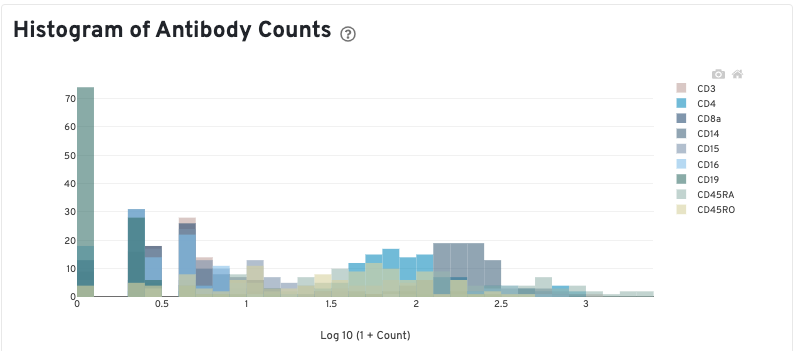
The Histogram of Antibody Counts can help to explore gene expression levels of the top 120 antibody marker genes across the library's population of cells. The x-axis shows UMI counts in the log10 scale (1=10 UMIs, 2=100 UMIs, etc.) and the y-axis shows the number of cells. To view this histogram for a specific subset of antibodies, select (or deselect) antibodies for display by clicking on the legend labels.
For information about the web summary for an Antibody Capture only analysis, go here.
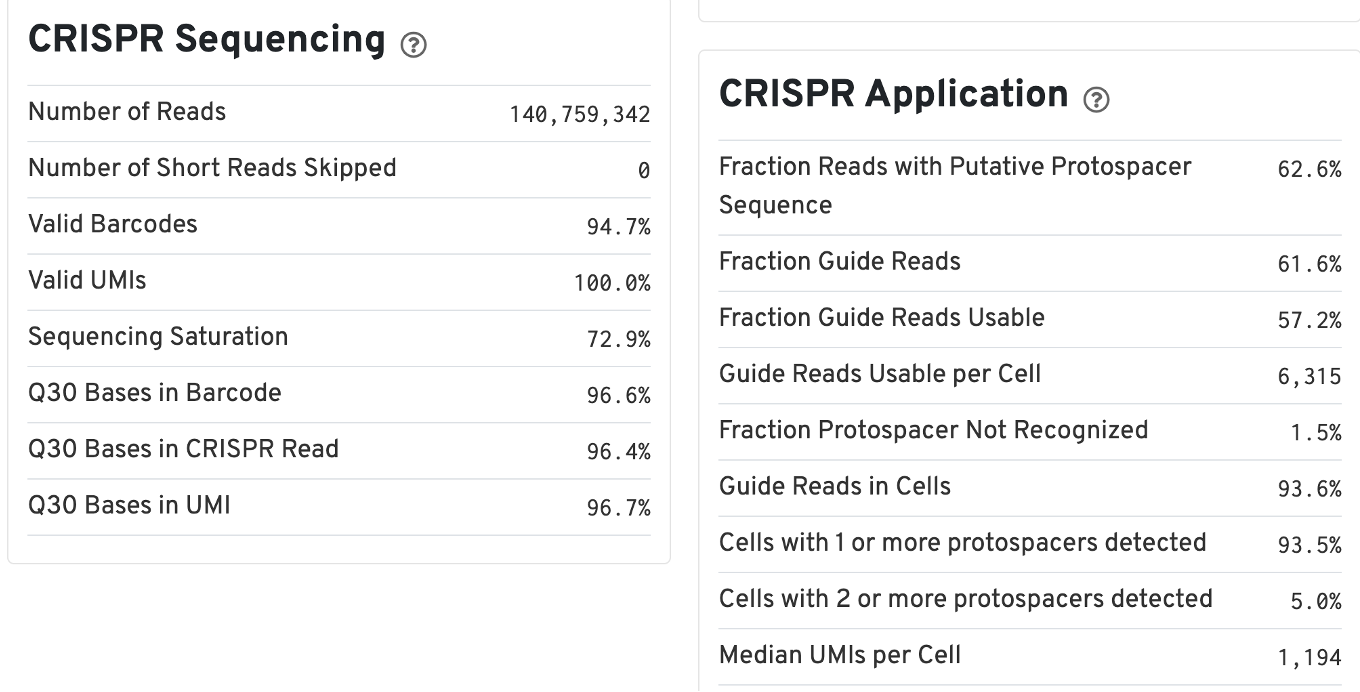
The web summary for Gene Expression and CRISPR Guide Capture libraries will have the same Summary and Gene Expression tabs. The Summary tab has two additional tables:
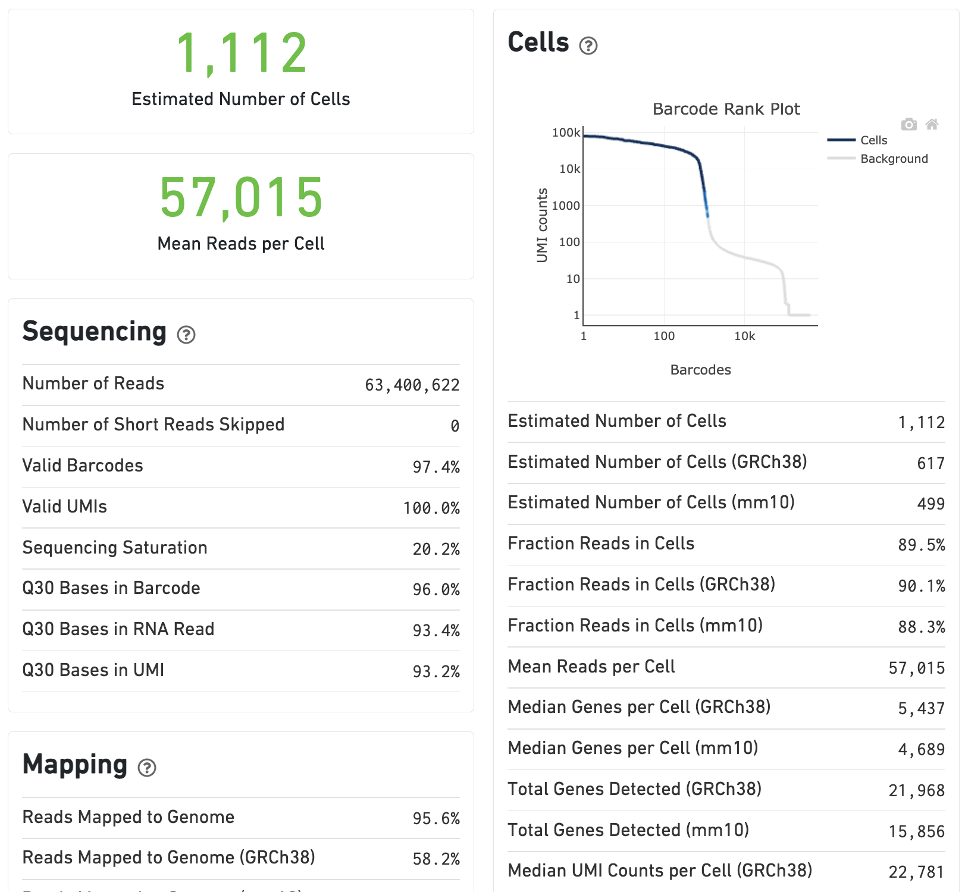
Single cell behavior can be demonstrated by experiments with a known mixture of cells from multiple species. For example, this human-mouse mixture experiment consists of approximately 500 human (HEK293T) cells and 500 mouse (3T3) cells in a 1:1 ratio.
The Summary tab will include metrics associated with mapping and cell counting for each reference transcriptome, in this case to human and mouse references. Click the ? at the top of each dashboard to display information about the metrics.
In the Gene Expression view, running the human-mouse mixture experiment results in the following GEM Partitions plot and metrics:
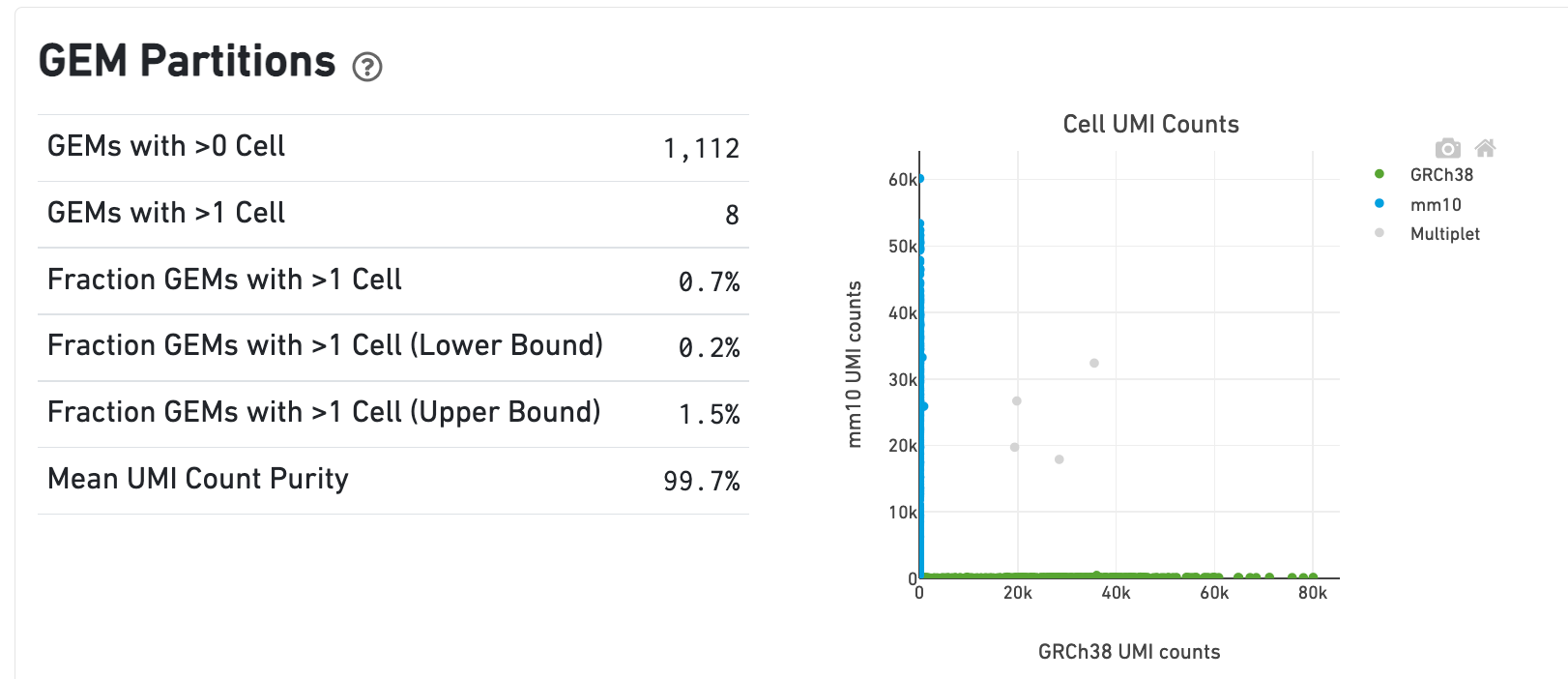
Each point represents a barcode. The gray points are inferred to be associated with GEM partitions that contain more than one cell because they contain a large number of molecules from human and mouse cells. The GEMs containing human-human and mouse-mouse multiplet cell combinations are not visible here, but their presence is inferred in the Fraction GEMs with >1 Cell metric shown next to the plot.
For Targeted Gene Expression experiments, the output includes modifications that highlight targeting performance. The following is an example of the Summary tab from a cellranger count targeted run from lymph node cells of a Hodgkin's Lymphoma patient.
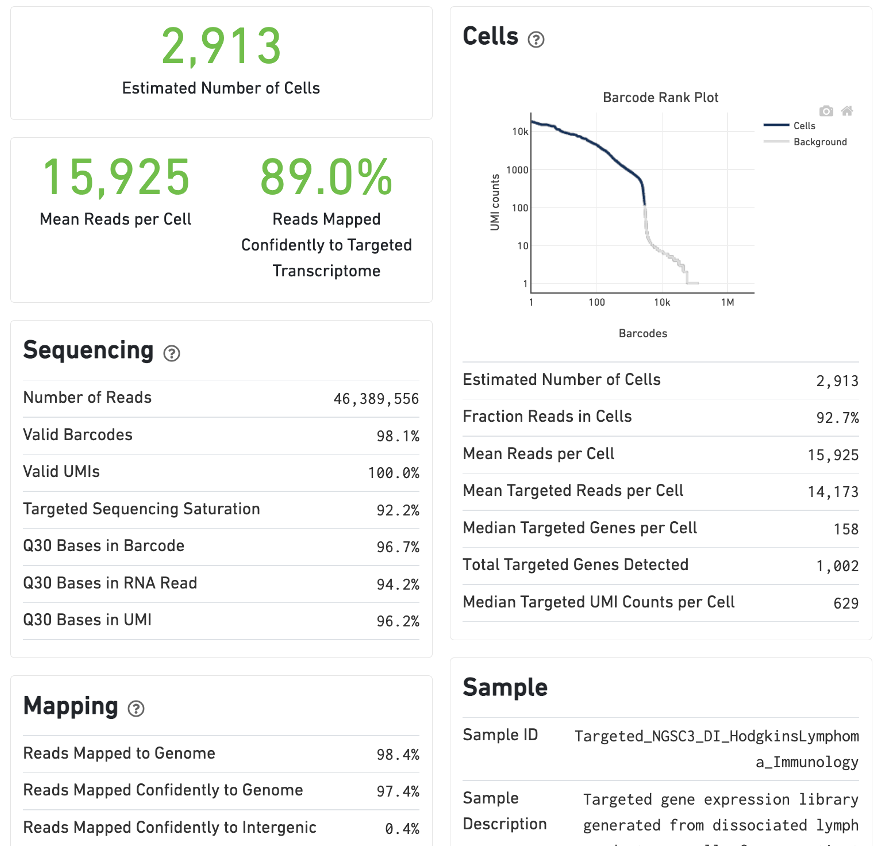
The Gene Expression tab includes a Targeted Enrichment dashboard with metrics that assess targeting performance. On the right is a scatterplot of read counts per gene versus UMI counts per gene. A successful targeted experiment where targeted genes are well-enriched looks like the one below, with targeted genes (blue) cleanly separated from non-targeted genes (gray). See Targeted Gene Expression Algorithms for more information on how gene enrichments are computed.
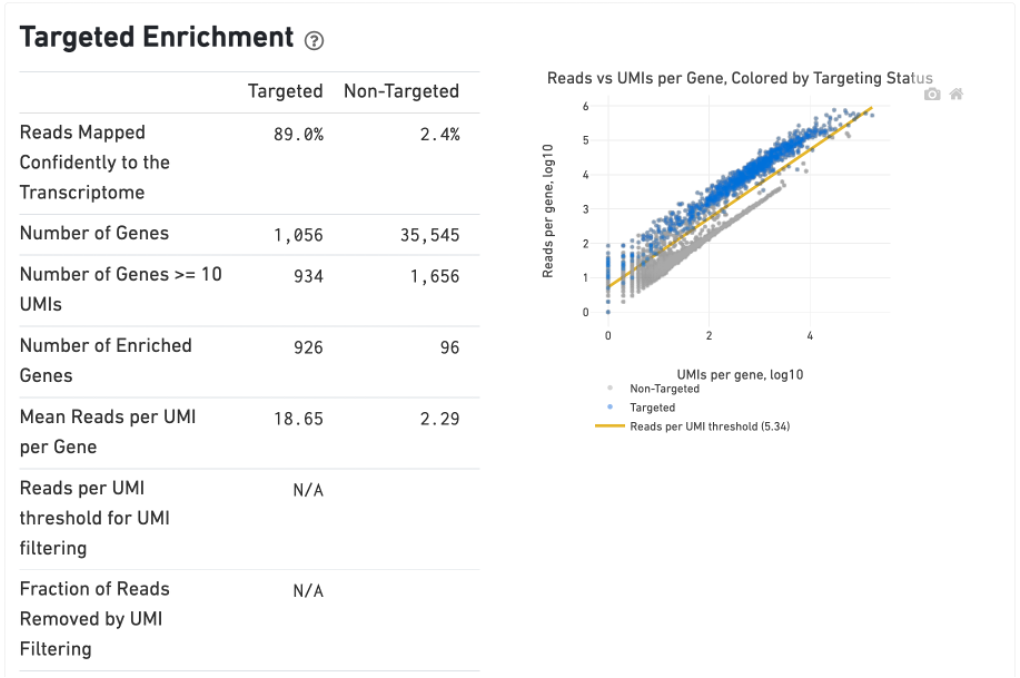
The remainder of the Gene Expression tab is very similar to that of a regular gene expression analysis, but focuses on targeted genes (Sequencing Saturation, Median Genes per Cell); all secondary analysis (t-SNE, differential expression) is done using only targeted genes.
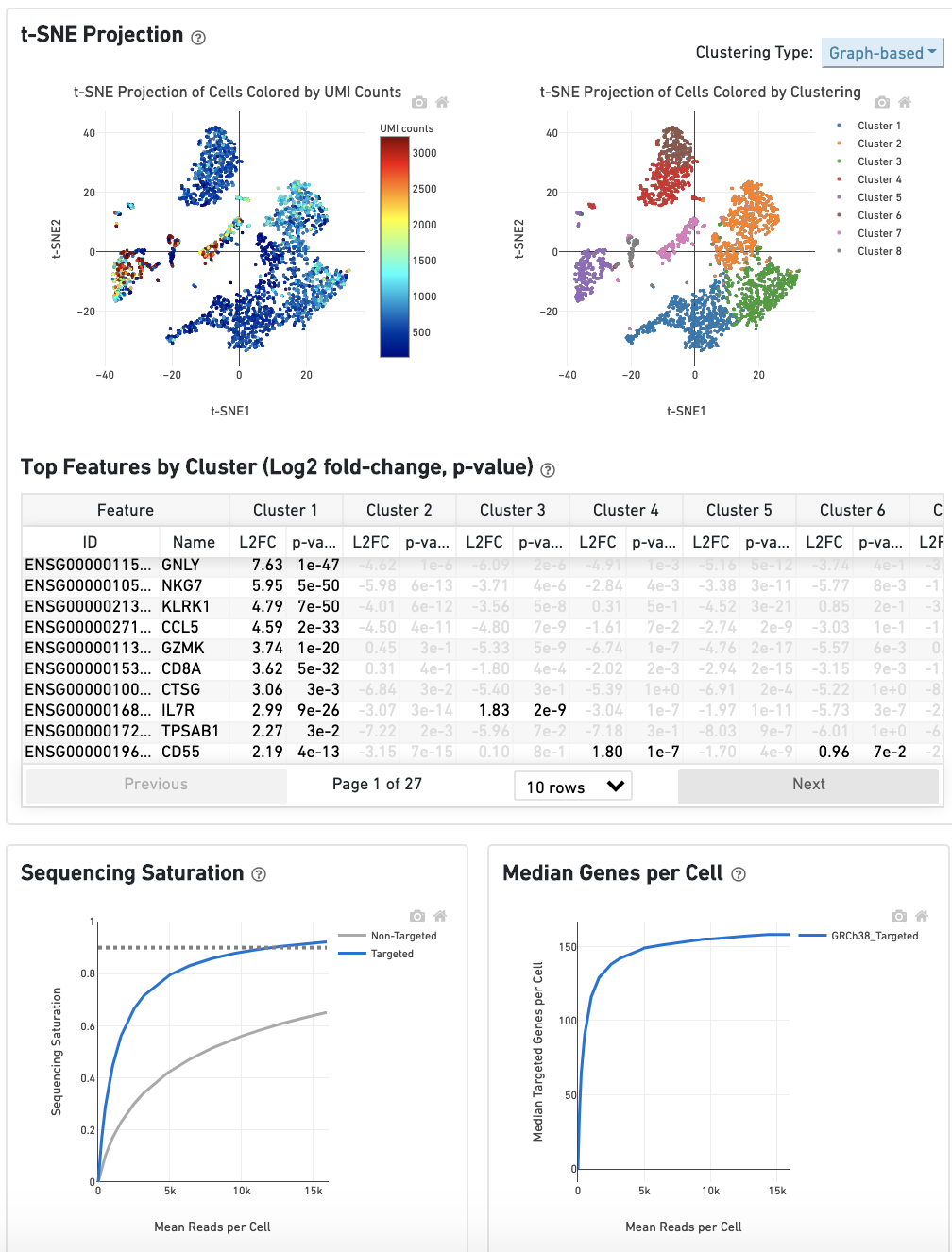
Do you have questions or feedback about this documentation? Please contact support@10xgenomics.com.