Cell Ranger6.1, printed on 03/30/2025
In this section, you will learn:
| Coming Soon: the next release of Cell Ranger will count intronic reads by default for whole transcriptome gene expression data. For more information, see our recommendation on including introns for gene expression analysis page. |
The 5' Chromium Single Cell Immune Profiling Solution with Feature Barcode technology enables simultaneous profiling of V(D)J repertoire, cell surface protein, antigen specificity, and gene expression (GEX) data. The cellranger multi pipeline analyzes these multiple library types together, enabling more consistent cell calling between the V(D)J and GEX data.
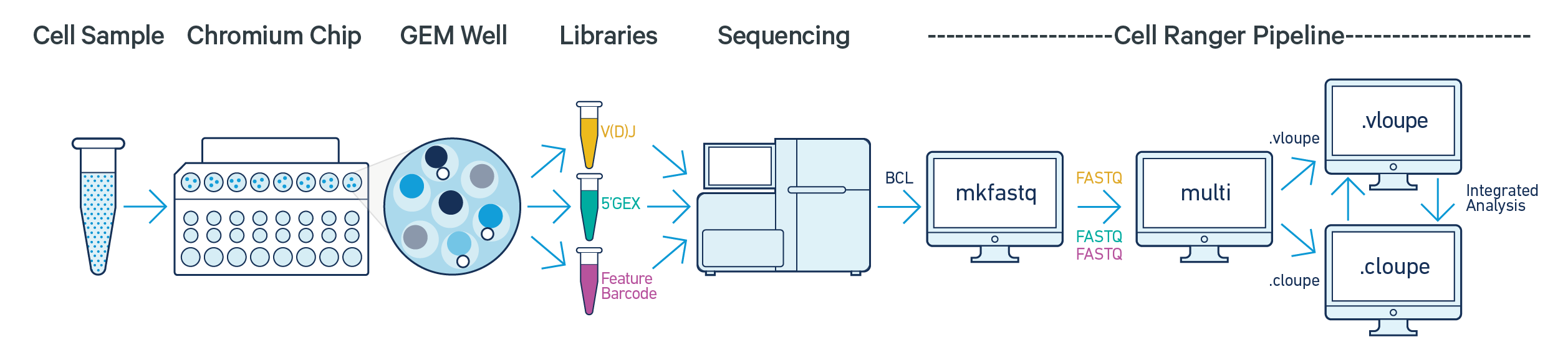
cellranger multi takes FASTQ files from cellranger mkfastq or bcl2fastq for any combination of 5' Gene Expression, Feature Barcode (cell surface protein or antigen), and V(D)J libraries from a single GEM well. It performs alignment, filtering, barcode counting, and UMI counting on the Gene Expression and/or Feature Barcode libraries. It also performs sequence assembly and paired clonotype calling on the V(D)J libraries. Additionally, the cell calls provided by the gene expression data are used to improve the cell calls from the V(D)J library. cellranger multi is the recommended pipeline for processing FASTQ files generated using the 5' the Immune Profiling Solution with Feature Barcode technology.
Pipeline recommendation depends on the combination of libraries being analyzed. This table summarizes all library combinations and corresponding pipeline recommendations:
| Use multi? | |||
|---|---|---|---|
| Recommended. Improves final cell calls in V(D)J library. Learn more. | |||
| Recommended. Improves final cell calls in V(D)J library. Learn more. | |||
| Optional. Alternative pipeline: count + vdj | |||
| Optional. Alternative pipeline: vdj | |||
| Optional. Alternative pipeline: count | |||
| Optional. Alternative pipeline: count | |||
| Optional. Alternative pipeline: count |
The cellranger multi pipeline improves cell calls in the V(D)J dataset by discarding any cells that were not also called in the corresponding 5' GEX dataset. By assigning cells that are called in the V(D)J results but not in the 5' GEX results as background GEMs in the V(D)J data, the cellranger multi pipeline mitigates any overcalling issues that may arise in BCR and TCR data. This improved cell calling is only possible when both 5' GEX and V(D)J libraries were sequenced from the same sample.
As shown in the image below, final V(D)J cell calls (green intersection area) exclude cells that were only called by the vdj pipeline (yellow region).
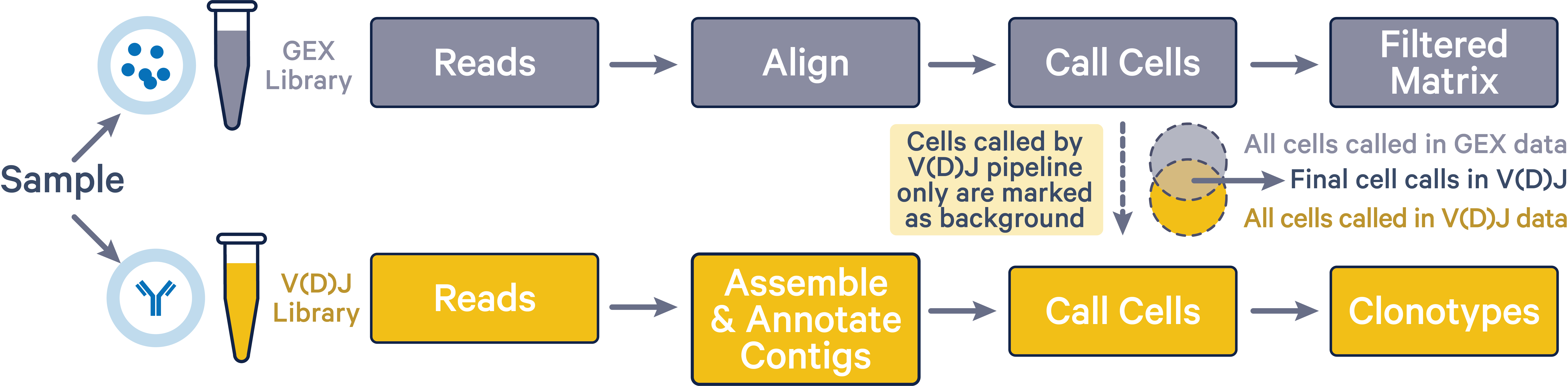
The 5' GEX cell calls are not affected by the cellranger multi pipeline. The gene expression library is representative of the entire pool of poly-adenylated mRNA transcripts captured within each partition (droplet). The TCR or BCR transcripts in the gene expression library are then selectively amplified to create the V(D)J library. Therefore, the gene expression library has more power to detect partitions containing cells compared to the V(D)J library. If the cellranger multi pipeline is run with both 5' GEX and V(D)J data, then barcodes which are not called as cells in the 5' GEX data will be deleted from the V(D)J cell set.
The cellranger multi pipeline takes FASTQ files as input. If you do not have FASTQ files for your V(D)J, single cell gene expression and/or Feature Barcode libraries, follow the instructions for running cellranger mkfastq to generate FASTQ files.
To simultaneously generate single cell feature counts, V(D)J sequences and annotations for a single library, run cellranger multi with the following arguments:
| Argument | Description |
|---|---|
--id | A unique run ID string: e.g. sample345 that is also the output folder name. Cannot be more than 64 characters. |
--csv | Path to multi config CSV file enumerating input libraries and analysis parameters. |
The multi config CSV contains both the library definitions and experiment configuration variables. It is composed of up to four sections: [gene-expression], [feature], [vdj], and [libraries]. The [gene-expression], [feature], and [vdj] sections have at most two columns, and are responsible for configuring their respective portions of the experiment. The [libraries] section specifies where input FASTQ files may be found. A customizable template for a multi config CSV can be downloaded here, and example multi config CSVs can be downloaded from public datasets. Example formats for two product configurations are below.
Starting with Cell Ranger 6.1, it is generally recommended to specify the estimated number of cells using expect-cells, a parameter for automated cell calling that specifies the expected number of recovered cells. Read more about the cell-calling algorithm on the 3' Single Cell Gene Expression page. Automated cell calling can be overridden with the force-cells option that bypasses the cell detection algorithm by drawing a hard cut off equal to the specified number barcodes with highest UMIs.
|
For help on how to configure the [libraries] section to target a particular set of FASTQs, consult Specifying Input FASTQ Files for cellranger multi.
|
After determining the input arguments, run cellranger multi. Remember to replace the bits of code in red with your sample id and csv file path:
mkdir /home/jdoe/runs cd /home/jdoe/runs cellranger multi --id=sample345 --csv=/home/jdoe/sample345.csv
Following a series of checks to validate input arguments, cellranger multi pipeline stages will begin to run:
Martian Runtime - v4.0.6 Running preflight checks (please wait)... ...
By default, cellranger will use all of the cores available on your
system to execute pipeline stages. You can specify a different number of cores
to use with the --localcores option; for example, --localcores=16
will limit cellranger to using up to sixteen cores at once. Similarly,
--localmem will restrict the amount of memory (in GB) used by
cellranger.
The pipeline will create a new folder named with the run ID you specified using the --id argument (e.g. /home/jdoe/runs/sample345) for its output. If this folder already exists, cellranger will assume it is an existing pipestance and attempt to resume running it. If you wish to start the run over, delete the output folder (sample345/ in this example) and rerun the pipeline.
A successful cellranger multi run should conclude with a message similar to this:
Waiting 6 seconds for UI to do final refresh. Pipestance completed successfully! yyyy-mm-dd hh:mm:ss Shutting down. Saving pipestance info to "tiny/tiny.mri.tgz"
To learn more about the output files generated, refer to the Outputs for multi section under Understanding Outputs.
Here are the example multi config CSVs for two library combinations. Make sure to replace /path/to with the actual full path to your data, and edit any text in red according to the experiment's sample/library/file names.
| Libraries | Multi config CSV |
 See example dataset |
[vdj] reference,/path/to/vdj_reference |
 See example dataset |
[gene-expression] reference,/path/to/transcriptome expect-cells,enter expected number of recovered cells include-introns,true |
 See example dataset |
[gene-expression] reference,/path/to/transcriptome expect-cells,enter expected number of recovered cells include-introns,true |
The cellranger multi pipeline supports downsampling the reads by specifying a rate between 0 and 1 independently for each library. It also allows trimming the reads to a fixed length, which is not supported in the cellranger vdj pipeline.
The option to run denovo without V(D)J reference (--denovo) is not supported in cellranger multi. This option is available in cellranger vdj
Next, you may wish to: