Cell Ranger2.0, printed on 04/27/2025
Cell Ranger for V(D)J is a set of analysis pipelines that process Chromium single cell 5’ RNA-seq output to assemble, quantify, and annotate paired V(D)J transcript sequences.
Cell Ranger for V(D)J includes two main pipelines:
cellranger mkfastq wraps Illumina's bcl2fastq to correctly demultiplex Chromium-prepared sequencing samples and to convert barcode and read data to FASTQ files.
cellranger vdj takes FASTQ files from cellranger mkfastq and performs V(D)J sequence assembly and paired clonotype calling. It uses the Chromium cellular barcodes and UMIs to assemble V(D)J transcripts cell-by-cell. cellranger can take input from multiple sequencing runs on the same library and/or from multiple libraries from the same sample.
Output is delivered in standard BAM, CSV, FASTA, FASTQ, JSON and HTML formats that are augmented with cellular and clonotype-specific information.
Throughout the documentation, you will see references to samples, libraries and sequencing runs. We define these as follows:
The relationship between these terms can be complex:
The Cell Ranger workflow always starts with running cellranger mkfastq on each flowcell directory, as described in Generating FASTQs. The subsequent steps vary depending on how many samples, libraries and flowcells you have. We will describe them in order of increasing complexity:
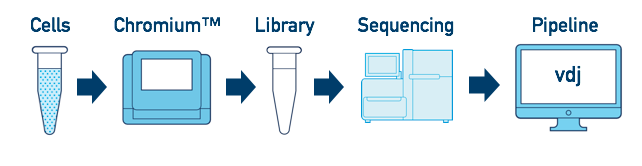
This is the most basic case. You have a single biological sample, which was prepared into a single library, and then sequenced on a single flowcell. Assuming the FASTQs have been generated with cellranger mkfastq, you just need to run cellranger vdj as described in V(D)J T Cell Analysis.
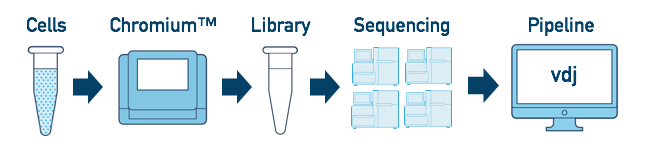
If you have a library which was sequenced across multiple flowcells, you can pool the reads from both sequencing runs. Follow the steps in Multi-Flowcell Samples to combine them in a single cellranger vdj run.
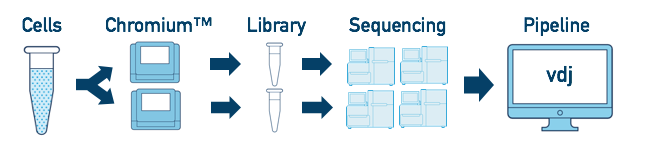
If you have a single sample which was run on multiple chip channels, producing multiple libraries, you can analyze all of the libraries at once. The workflow is similar to the "Multiple Flowcell" workflow above. Follow the steps in Multi-Flowcell Samples to combine them in a single cellranger vdj run.