Cell Ranger ATAC2.1, printed on 03/29/2025
Cluster mode is one of three primary ways of running Cell Ranger ATAC. To learn about the other approaches, please go to the computing options page.
Cell Ranger ATAC can be run in cluster mode, using SGE or LSF to run the stages on multiple nodes via batch scheduling. This allows highly parallelizable stages to utilize hundreds or thousands of cores concurrently, dramatically reducing time to solution.
| 10x Genomics does not officially support Slurm or Torque/PBS. Some customers have successfully used Cell Ranger ATAC with those job schedulers in cluster mode. However, it is unsupported and may require trial and error attempts. |
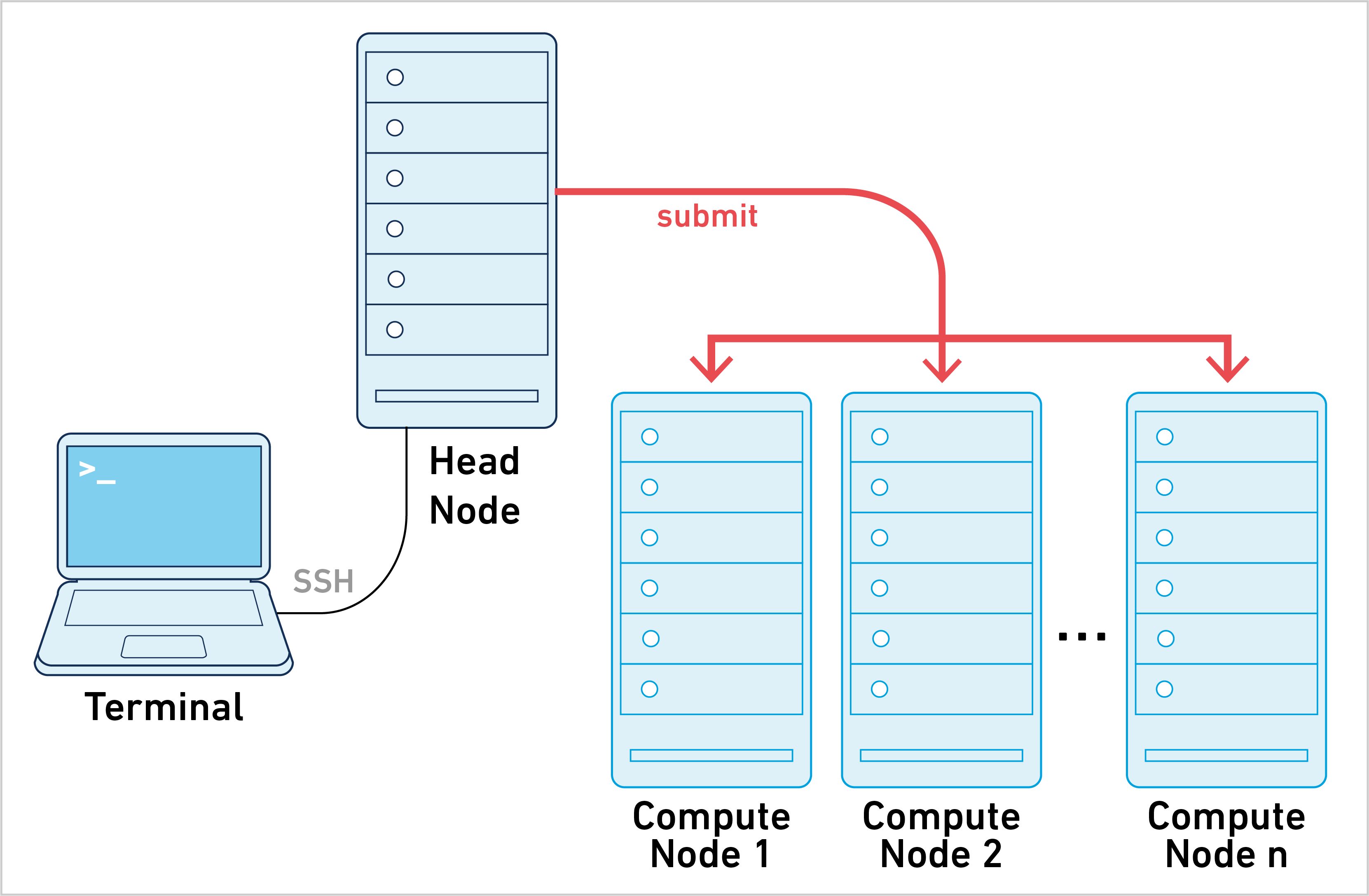
Running pipelines in cluster mode requires the following:
/opt/cellranger-atac-2.1.0
or
/net/apps/cellranger-atac-2.1.0Installing the Cell Ranger ATAC software on a cluster is identical to installation on a local server. After confirming that the cellranger-atac commands can run in single server mode, configure the job submission template that Cell Ranger ATAC uses to submit jobs to the cluster. Assuming Cell Ranger ATAC is installed to /opt/cellranger-atac-2.1.0, the process is as follows.
Step 1. Navigate to the /external/martian/jobmanagers directory which contains example jobmanager templates.
$ cd /opt/cellranger-atac-2.1.0/external/martian/jobmanagers $ ls bsub.template.example config.json sge.template.example
Step 2. Make a copy of the cluster's example template (SGE or LSF) to either sge.template or lsf.template in the jobmanagers/ directory.
$ cp -v sge.template.example sge.template `sge.template.example' -> `sge.template' $ ls bsub.template.example config.json sge.template sge.template.example
The job submission templates contain a number of special variables that are substituted by the Martian runtime when each stage is submitted. Specifically, the following variables are expanded when a pipeline is submitting jobs to the cluster:
| Variable | Must be present? | Description |
|---|---|---|
__MRO_JOB_NAME__ |
Yes | Job name composed of the sample ID and stage being executed |
__MRO_THREADS__ |
No | Number of threads required by the stage |
__MRO_MEM_GB____MRO_MEM_MB__ |
No | Amount of memory (in GB or MB) required by the stage |
__MRO_MEM_GB_PER_THREAD____MRO_MEM_MB_PER_THREAD__ |
No | Amount of memory (in GB or MB) required per thread in multi-threaded stages. |
__MRO_STDOUT____MRO_STDERR__ |
Yes | Paths to the _stdout and _stderr metadata files for the stage |
__MRO_CMD__ |
Yes | Bourne shell command to run the stage code |
It is critical that the special variables listed as required are present in the final template you create. For more information on how the template should appear for a cluster, consult your cluster administrator or help desk.
Depending on which job scheduler you have, please select a tab below.
<pe_name> within the example template to reflect the name of the cluster's multithreaded parallel environment. To view a list of the cluster's parallel environments, use the qconf -spl command.
The most common modifications to the job submission template include adding additional lines to specify:
#$ -q smith.q#$ -A smith_lab#BSUB -q smith_queue#BSUB -P smith_labTo run a Cell Ranger ATAC pipeline in cluster mode, add the --jobmode=sge
or --jobmode=lsf command-line option when using the cellranger-atac commands.
It is also possible to use --jobmode=<PATH>, where <PATH> is the full path to the cluster template file.
To validate that cluster mode is properly configured, follow the same
validation instructions given for cellranger-atac in the Installation
page, but add --jobmode=sge or --jobmode=lsf.
$ cellranger-atac mkfastq --run=./tiny-bcl --samplesheet=./tiny-sheet.csv --jobmode=sge Martian Runtime - 2.1.0 Running preflight checks (please wait)... ...
Once the preflight checks are finished, check the job queue to see stages queuing up:
$ qstat job-ID prior name user state submit/start at queue slots ja-task-ID ----------------------------------------------------------------------------------------------------------------- 8675309 0.56000 ID.HAWT7AD jdoe qw 09/13/2016 12:00:00 all.q@cluster.university.edu 1 8675310 0.55500 ID.HAWT7AD jdoe qw 09/13/2016 12:00:00 all.q@cluster.university.edu 1
In the event of a pipeline failure, an error message is displayed.
[error] Pipestance failed. Please see log at: HAWT7ADXX/MAKE_FASTQS_CS/MAKE_FASTQS/MAKE_FASTQS_PREFLIGHT/fork0/chnk0/_errors Saving diagnostics to HAWT7ADXX/HAWT7ADXX.debug.tgz For assistance, upload this file to 10x by running: uploadto10x <your_email> HAWT7ADXX/HAWT7ADXX.debug.tgz
The _errors file contains a jobcmd error:
$ cat HAWT7ADXX/MAKE_FASTQS_CS/MAKE_FASTQS/MAKE_FASTQS_PREFLIGHT/fork0/chnk0/_errors jobcmd error: exit status 1
The most likely reason for this failure is an invalid job submission template. This occurs when the job submission via qsub or bsub commands failed.
The "tiny" dataset doesn't stress the cluster, so it's worthwhile to follow up with a more realistic test using one of our sample datasets.
There are two subtle variants of running cellranger-atac pipelines in cluster mode, each with their own pitfalls. Check with your cluster administrator to see which approach is compatible with your institution's setup.
--jobmode=sge on the head node. Cluster mode was originally designed for this use case. However, this approach leaves mrp and mrjob running on the head node for the duration of the pipeline, and some clusters impose time limits to prevent long running processes.--jobmode=sge: With this approach, mrp and mrjob run on a cluster mode. However, the cluster must allow jobs to be submitted from a compute node to make this viable.When cellranger-atac is run in cluster mode, a single library analysis is partitioned into hundreds and potentially thousands of smaller jobs. The underlying Martian pipeline framework launches each stage job using the qsub or bsub commands when running on SGE or LSF cluster, respectively. As stage jobs are queued, launched, and completed, the pipeline framework tracks their status using the metadata files that each stage maintains in the pipeline output directory.
Like single server pipelines, cluster-mode pipelines can be restarted after failure. They maintain the same order of execution for dependent stages of the pipeline. All executed stage code is identical to single server mode, and the quantitative results are identical to the limit of each stage's reproducibility.
| Cluster-mode pipelines that are stopped (either by you or due to a stage failure) do not delete pending stages that have already been submitted to the cluster queue. As a result, some pipeline stages may continue to execute after the cellranger-atac commands have exited. |
In addition, the Cell Ranger ATAC UI can still be used with cluster mode. Because the Martian pipeline framework runs on the node from which the command was issued, the UI will also run from that node.
Stages in the Cell Ranger ATAC pipelines each request a specific number of cores and memory to aid with resource management. These values are used to prevent oversubscription of the computing system when running pipelines in single server mode. The way CPU and memory requests are handled in cluster mode is defined by the following:
__MRO_THREADS__ and
__MRO_MEM_GB__
variables are used within the job template.Depending on which job scheduler you have, select a tab below.
mem_free resource natively,
although your cluster may have another mechanism for requesting memory. To pass each stage's
memory request through to SGE, add an additional line to your
sge.template that requests mem_free,
h_vmem, h_rss,
or the custom memory resource defined by your cluster:
$ cat sge.template #$ -N __MRO_JOB_NAME__ #$ -V #$ -pe threads __MRO_THREADS__ #$ -l mem_free=__MRO_MEM_GB__G #$ -cwd #$ -o __MRO_STDOUT__ #$ -e __MRO_STDERR__ __MRO_CMD__
In the above example, the trailing G in the highlighted
__MRO_MEM_GB__G is required by SGE to denote that mem_free is being expressed
in GB units.
|
Note that the h_vmem (virtual memory) and mem_free/h_rss (physical
memory) represent two different quantities, and that Cell Ranger ATAC stages'
__MRO_MEM_GB__ requests are expressed as physical memory. Using h_vmem in your job template may cause certain stages
to be unduly killed if their virtual memory consumption is substantially larger
than their physical memory consumption. It follows that we do not recommend
using h_vmem.
| If you do use h_vmem in a template, it is recommended that you use the __MRO_MEM_GB_PER_THREAD__ or __MRO_MEM_MB_PER_THREAD__ variables instead of __MRO_MEM_GB__ and __MRO_MEM_MB__. To determine memory limits for a multicore job, SGE will multiply the number of threads by the value in h_vmem. The __MRO_MEM_GB__ and __MRO_MEM_MB__ variables already reflect the sum amount of memory across all threads needed to run the job. Using those variables as h_vmem will inflate the memory required for multi-threaded jobs. |
-M and
-R [mem=...] options, but these requests generally must
be expressed in MB, not GB. As such, your LSF job template should use the
__MRO_MEM_MB__ variable rather than
__MRO_MEM_GB__. For example,
$ cat bsub.template #BSUB -J __MRO_JOB_NAME__ #BSUB -n __MRO_THREADS__ #BSUB -o __MRO_STDOUT__ #BSUB -e __MRO_STDERR__ #BSUB -R "rusage[mem=__MRO_MEM_MB__]" #BSUB -R span[hosts=1] __MRO_CMD__
For clusters whose job managers do not support memory requests, it is possible
to request memory in the form of cores via the --mempercore
command-line option. This option scales up the number of threads requested
via the __MRO_THREADS__ variable according to how much memory
a stage requires.
For example, given a cluster with nodes that have 16 cores and 128 GB of memory (8 GB per core), the following pipeline invocation command
$ cellranger-atac mkfastq --run=./tiny-bcl --samplesheet=./tiny-sheet.csv --jobmode=sge --mempercore=8
will issue the following resource requests:
__MRO_THREADS__ of 1 to the job template.__MRO_THREADS__ of 2 to the job template because (12 GB) / (8 GB/core) = 2 cores.__MRO_THREADS__ of 2 to the job template.__MRO_THREADS__ of 5 to the job template because (40 GB) / (8 GB/core) = 5 cores.As the final bullet point illustrates, this mode can result in wasted CPU cycles and is only provided for clusters that cannot allocate memory as an independent resource.
Every cluster configuration is different. If you are unsure of how your cluster resource management is configured, contact your cluster administrator or help desk.
Some Cell Ranger ATAC pipeline stages are divided into hundreds of jobs. By default, the rate at which these jobs are submitted to the cluster is throttled to at most 64 at a time and at least 100ms between each submission to avoid running into limits on clusters which impose quotas on the total number of pending jobs a user can submit.
If your cluster does not have these limits or is not shared with other users, you can control
how the Martian pipeline runner sends job submissions to the cluster by using the
--maxjobs and --jobinterval parameters.
To increase the cap on the number of concurrent jobs to 200, use the --maxjobs parameter:
$ cellranger-atac count --id=sample ... --jobmode=sge --maxjobs=200
You can also change the rate limit on how often the Martian pipeline runner sends submissions to
the cluster. To add a five-second pause between job submissions, use the --jobinterval parameter:
$ cellranger-atac count --id=sample ... --jobmode=sge --jobinterval=5000
The job interval parameter is in milliseconds. The minimum allowable value is 1.