Long Ranger2.2, printed on 03/29/2025
| Analysis software for 10x Genomics linked read products is no longer supported. Raw data processing pipelines and visualization tools are available for download and can be used for analyzing legacy data from 10x Genomics kits in accordance with our end user licensing agreement without support. |
We recommend running longranger with GATK for more accurate calling of SNPs and indels. You must already have a working version of GATK installed on your system. Support is also provided for using a bundled build of Freebayes. Long Ranger uses GATK's HaplotypeCaller mode. See the GATK installation instructions
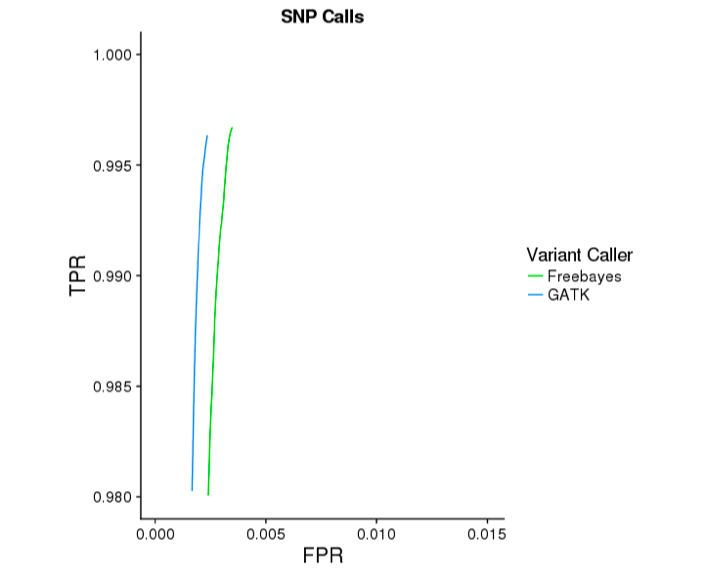
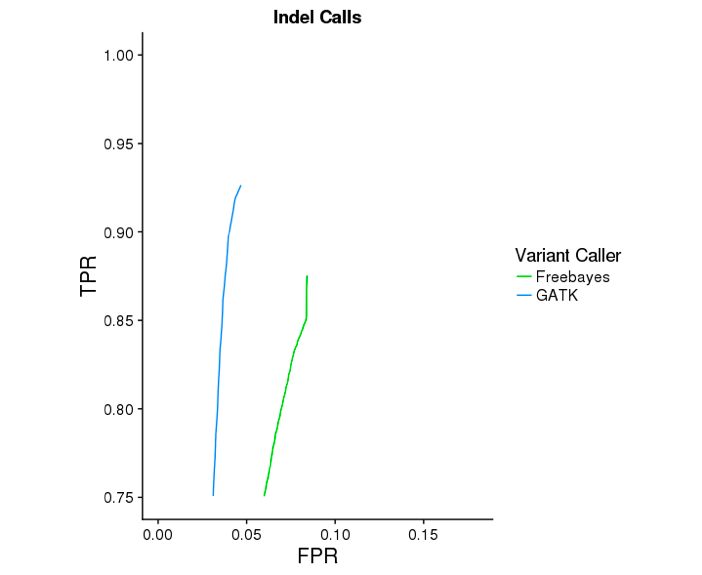
The GATK mode of Long Ranger yields substantially more accurate indel calls. Below are ROC curves reported by hap.py for 30x WGS Chromium data on NA12878, against the Genome In a Bottle v3.2.2 reference variants. GATK improves indel sensitivity by ~5 percentage points, while improving specificty.
| Long Ranger Version | GATK Version Supported |
|---|---|
| 2.2 and later | 3.3-4.0, excluding 3.6 |
| 2.1 and earlier | 3.3-3.8, excluding 3.6 |
| Changes to GATK occurring after version 4.0.3 have introduced an incompatibility with Long Ranger. We will release an update to Long Ranger to account for this, but in the meantime the maximum version level for GATK that will work is 4.0.3. You can download GATK 4.0.3 from here. |
To use GATK, please note the following:
longranger mkref will instruct you on how to create a genome.dict file. You will need this file to use GATK.You can instruct longranger to use GATK as its variant caller by passing the --vcmode=gatk:/path/to/GenomeAnalysisTK.jar parameter. For example, if your GenomeAnalysisTK.jar jarfile is located in /usr/local/gatk/GenomeAnalysisTK.jar,
$ longranger wgs --id=sample345 \ --sex=female \ --fastqs=/home/jdoe/runs/HAWT7ADXX/outs/fastq_path \ --indices=SI-GA-A1 \ --reference=/opt/refdata-hg19-2.1.0 \ --targets=/home/jdoe/runs/agilent_exome.bed \ --vcmode=gatk:/usr/local/gatk/GenomeAnalysisTK.jar \ --uiport=3600
Aside from the stage where SNPs and indels are called, Long Ranger will behave the same whether using GATK or Freebayes.
| The path to the GATK JAR must be an absolute path (e.g., /home/jdoe/bin/GenomeAnalysisTK.jar, not just bin/GenomeAnalysisTK.jar). |