Space Ranger1.1, printed on 03/30/2025
10x recommends sequencing read 2 to a length of 90 bases.
Space Ranger 1.1 is less sensitive than previous versions to read length effects. In our experience, the rate at which reads map confidently to a gene, as well as the sensitivity of the resulting analysis, is consistent across a range of read lengths longer than 70 bp. The following graph shows the relationship between read mapping and read 2 length, after trimming, for six representative samples.
A full length cDNA construct is flanked by the 30 bp template switch oligo (TSO) sequence, AAGCAGTGGTATCAACGCAGAGTACATGGG, on the 5' end and poly-A on the 3' end. Some fraction of sequencing reads are expected to contain either or both of these sequences, depending on the fragment size distribution of the sequencing library. Reads derived from short RNA molecules are more likely to contain either or both TSO and poly-A sequence than longer RNA molecules.
Since the presence of non-template sequence in the form of either template switch oligo (TSO) or poly-A, low-complexity ends confound read mapping, TSO sequence is trimmed from the 5' end of read 2 and poly-A is trimmed from the 3' end prior to alignment. Trimming improves the sensitivity of the assay as well as the computational efficiency of the software pipeline.
Tags ts:i and pa:i in the output BAM files indicate the number of TSO nucleotides trimmed from the 5' end of read 2 and the number of poly-A nucleotides trimmed from the 3' end. The trimmed bases are present in the sequence of the BAM record and are soft clipped via the CIGAR string.
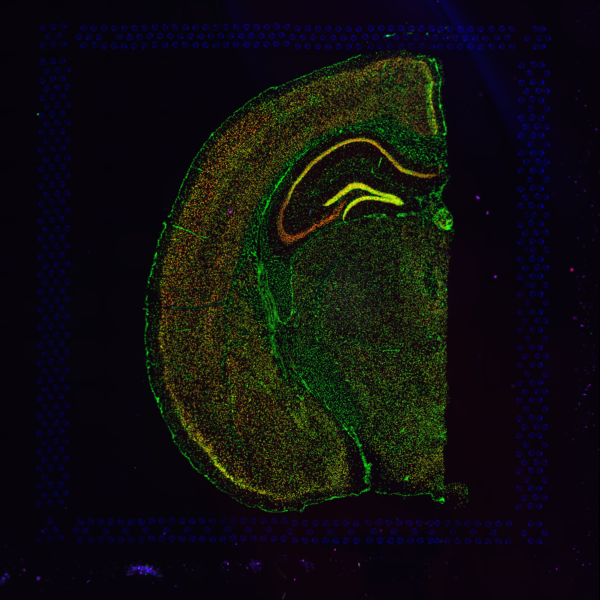
Space Ranger supports two styles of microscopy images: either a brightfield image stained with hematoxylin and eosin (H&E) or a fluorescence image comprising one or more acquisitions or channels (see below for examples of both, first row). The typical brightfield image is a transmission image, illuminated from below with white light and imaged from above. Dense areas of the sample, where tissue and cells lie, attenuate the transmitted light, producing an image that has a bright background and a dark or colored foreground. Fluorescence microscopy uses excitation with narrow band light and measures a signal emitted from either endogenous or added fluorophores, generating an image with a dark background and bright foreground. The fluorescence imaging demonstrated protocol for Visium is for immunofluorescence, where signal is generated by fluorophores conjugated to antibodies that bind to proteins of interest.
A fluorescence image comprises one or more grayscale intensity images (below, bottom row) and is often visualized as a single combined image with a different color assigned to each channel acquired (below, top-right). Each channel corresponds to a different fluorophore that binds to a different protein of interest. Space Ranger can take as input a brightfield image, one or more grayscale fluorescence channel images, or a color image made from combining and coloring fluroescence channel images in an external program such as ImageJ/Fiji or Photoshop.
 |
 |
| Brightfield Image | Fluorescence Image Colorized |
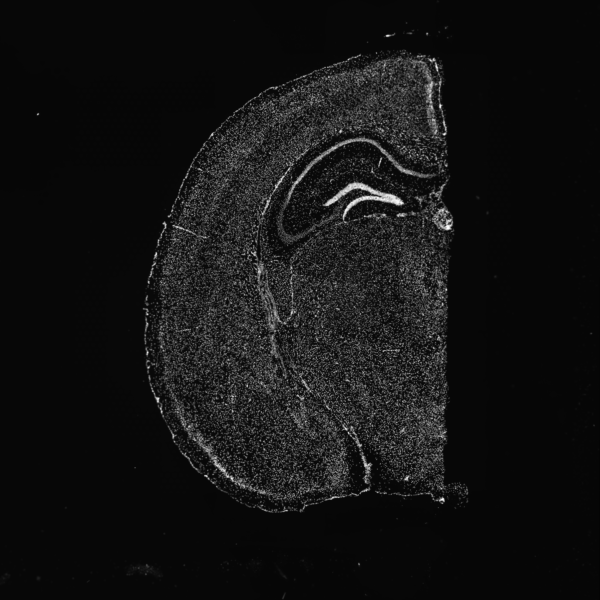 |
 |
 |
| Fluorescence Image, channel 1 | Fluorescence Image, channel 2 | Fluorescence Image, channel 3 |
Space Ranger requires images that are at least 2000 pixels in either dimension. In order to ensure uniform performance of the image processing in Space Ranger across a range of input resolutions, the input image is downsampled to be no larger than 2000 pixels in either dimension. Space Ranger's automatic processing will not perform well on smaller images and they are explicitly rejected from processing. Note that any downsampling performed internally in Space Ranger does not impact visualization in Loupe Browser, which always uses the full resolution input image.
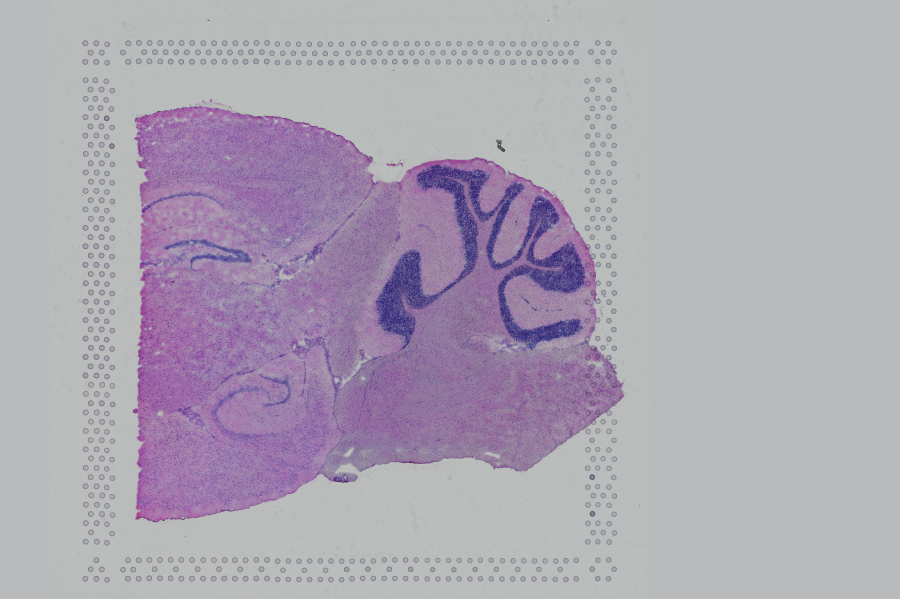
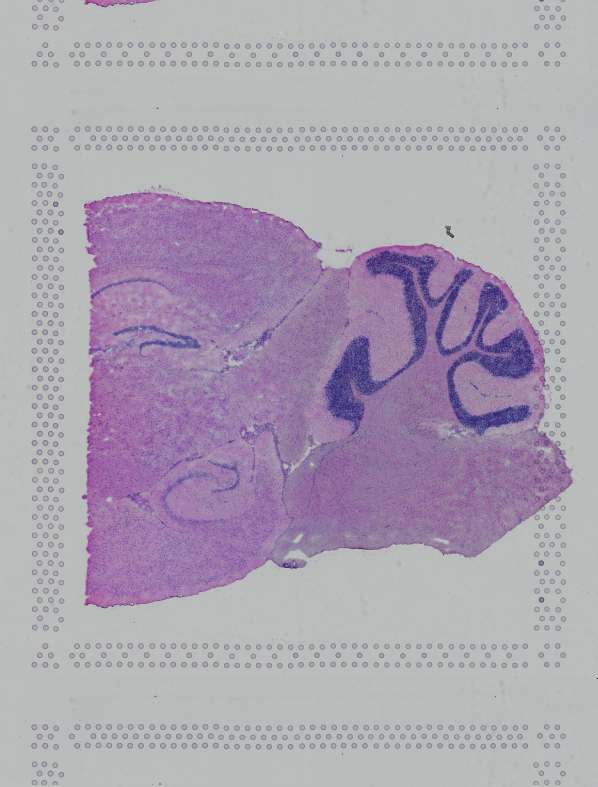
If the input image has a great deal of whitespace to the left or right of the capture area, or the image includes a portion of the capture area either above or below the area being analyzed, the capture area in the input image may be downsampled too much in order to accommodate the extra white space. If the fiducial alignment or tissue detection results are not as expected for such images, we recommend cropping the images in order to remove the extraneous image area outside of the fiducial border.
 |
 |
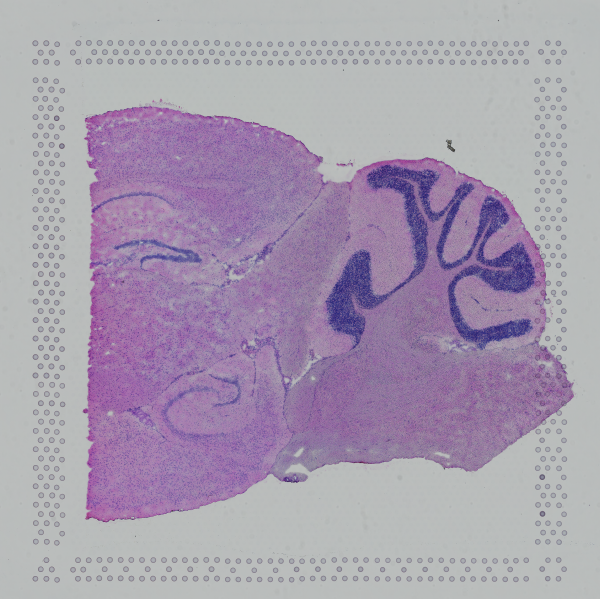 |
| Too Wide | Too Tall | Correct |
Images that are too tall or too wide can be easily cropped with freely available tools such as ImageJ/Fiji.
Brightfield microscopy input images for use with Visium must be 24-bit color TIFF, 16-bit grayscale TIFF, or JPEG.
In addition to these basic file type requirements for both Loupe Browser and Space Ranger, the automatic image processing pipeline in Space Ranger can be used for brightfield images that meet the additional restrictions outlined below. If these restrictions cannot be met, you can still process your data using the manual alignment and tissue selection process in Loupe Browser.
Images destined for automatic processing in Space Ranger should be oriented such that the hourglass fiducial marker is the in upper-left corner of the image. The image must be roughly axis-aligned, although slight rotations (e.g. less than 15 degrees) should be fine.
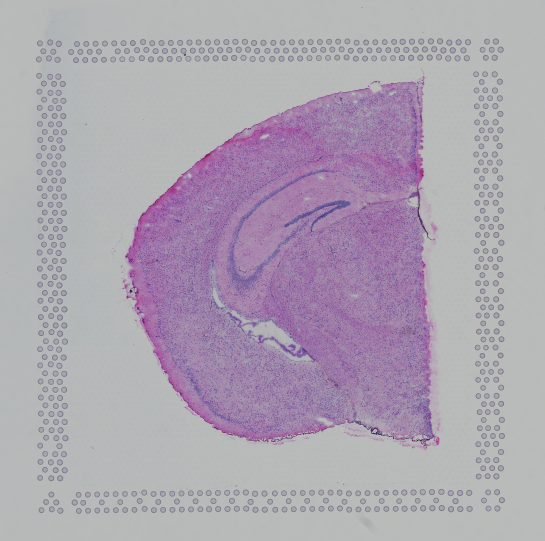 |
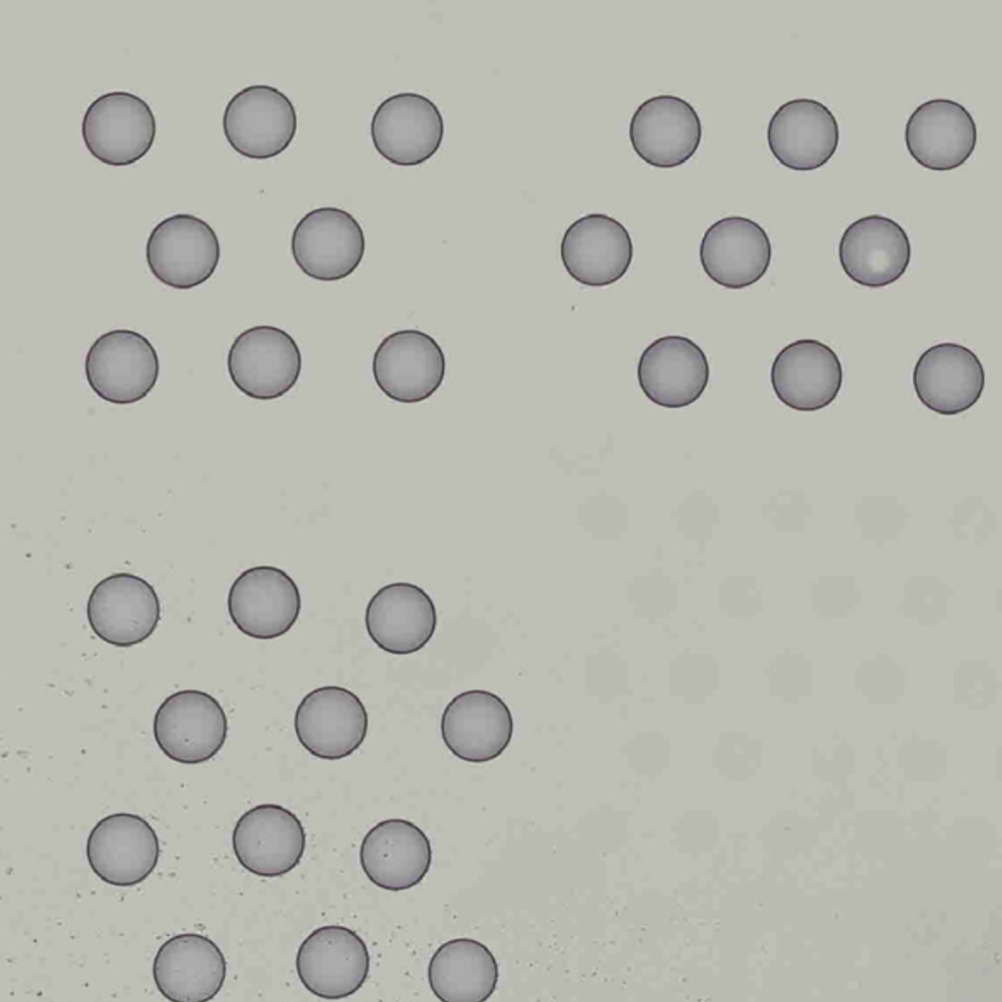 |
| Properly Oriented Slide Image | Corner with Hourglass Fiducial |
If the --reorient-images option is supplied to spaceranger count, the software will attempt to correct for images that are not in the expected orientation. This process is disabled by default as it is more likely to fail if tissue placement obscures the edges of the fiducial frame. If this process fails, the user can also correct for image rotation and mirroring either by using the manual alignment step in Loupe Browser or by correcting the image using freely available tools such as ImageJ/Fiji or Image Magick prior to processing with Space Ranger.
The automatic image processing in Space Ranger relies on the detection of spots in the fiducial frame. This process may perform poorly if the images are overexposed. Because the fiducial spots are slightly darker at the edges, very overexposed fiducials will appear as rings instead of solid circles. In general, poorly exposed images cannot be corrected after acquisition. If such images fail to process properly in the automatic pipeline, you can process such data through the manual alignment and tissue selection process in Loupe Browser prior to running Space Ranger.
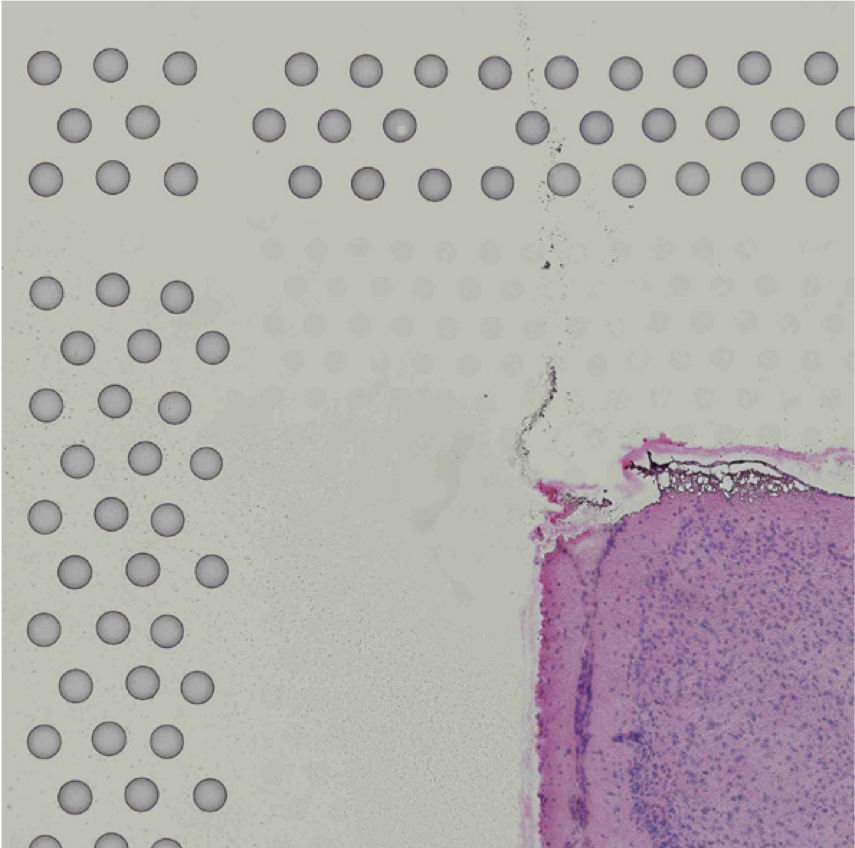 |
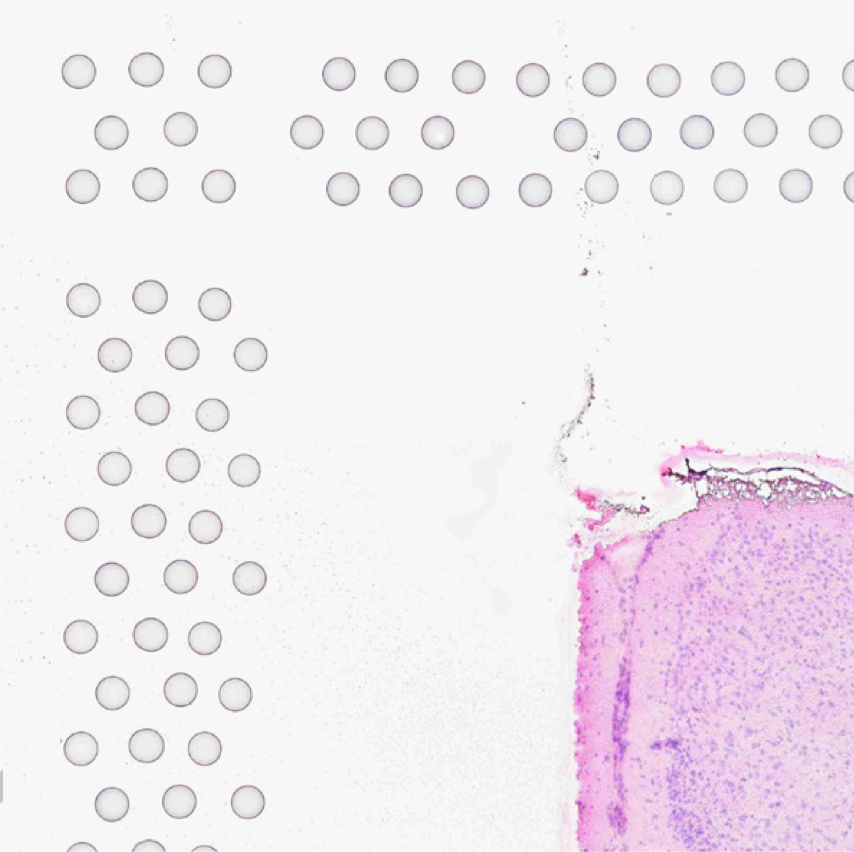 |
| Properly Exposed Slide Image | Over-Exposed Slide Image |
Fluorescence images can be provided to the pipeline either as individual grayscale acquisitions or as a single, combined, color image. There are three possible input modes:
One or more acquisitions stored in a single, multi-page TIFF file, where each page encodes an acquisition in greyscale at 8 or 16-bit depth. All the pages must be the same dimensions in pixels and the same bit depth.
One or more acquisitions each stored in individual, single-page TIFF files of JPEG files with the same restrictions as above. All files must be the same size and bit depth.
A color image where the user has colored and combined the individual acquisitions into a single image. This format can be specified as either a 24-bit color TIFF or a JPEG file. Note that with the options above, Loupe Browser will allow you to combine and color the images during visualization. This option is provided for users who wish to perform this function externally in software such as Photoshop or ImageJ/Fiji.
Space Ranger will not accept images in other formats at this time, but we're always open to suggestions for preferred image formats.
Important: Processing of fluorescence microscopy requires manual alignment and tissue area selection using manual alignment in Loupe Browser. The automatic registration and tissue detection analysis is not currently supported for fluorescence images.