Cell Ranger2.2, printed on 03/30/2025
With the Chromium Single Cell Immune Profiling Solution, it is now possible to obtain both the gene expression profile and T cell receptor and B cell immunoglobulin repertoires from the same input sample. Merging V(D)J and gene expression data will allow users to analyze the adaptive immune response inside complex tissue samples, observe the effects of immunotherapies, and investigate complex autoimmune, infectious, and other immune diseases.
You will need to download the latest versions of Cell Ranger, Loupe Cell Browser, and Loupe V(D)J Browser to get started.
What follows on this page is a high-level guide to conducting integrated V(D)J and gene expression experiments. You will find deeper information in the links within each section.
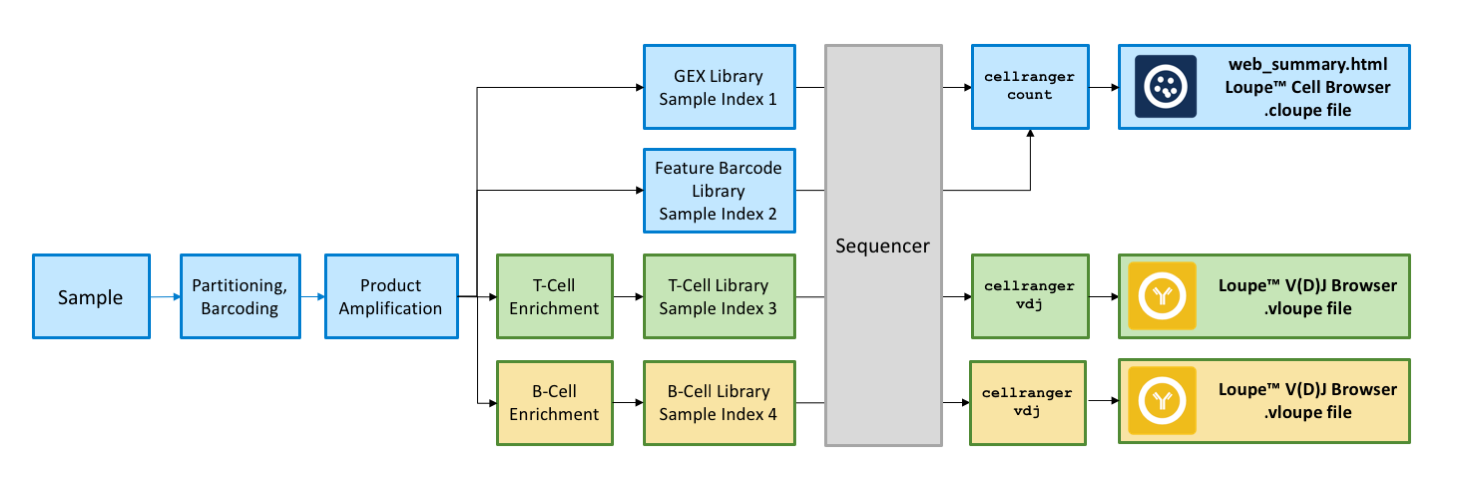
The graphic above shows an overview of the integrated 5′ V(D)J and Gene Expression workflow. From your input sample, you will create a single batch of barcoded droplets, create cDNA libraries for each cell, and amplify the cDNA without targeted enrichment. At that point, you will split the sample into two or more aliquots. The unenriched aliquot, shown in blue, is ready for sample indices and sequencing. To the aliquots marked for either T cell (green) or B cell (yellow) enrichment, you will add primers from the Chromium Human T Cell or B Cell Kits, perform a few cycles of additional amplification, and then prepare your sequencing libraries. At the end, you will have either two or three separate libraries from the same input sample, identified with separate I7 indices.
For the complete workflow, please consult the Chromium™ Single Cell V(D)J Reagent Kit User Guide.
You will treat your 5′ gene expression and enriched aliquots as separate libraries on the sequencer. Both the unenriched and enriched libraries can be pooled on the same sequencing run, though it may be more cost-effective to run the gene expression library separately. The configurations are:
The libraries are compatible with Illumina MiSeq, HiSeq 2500 (Rapid Run/High Output), HiSeq 3000/4000, NovaSeq, and NextSeq sequencers. For more assay and sequencing-specific information, consult the Single Cell V(D)J Sequencing page.
For the remainder of this page, we will suppose a sample sheet was used that yielded FASTQs
with sample name prefixes SampleGEX, SampleT and SampleB.
To process the three different libraries, you will need to run three different instances of
the new Cell Ranger 2.1 pipelines. For the gene expression SampleGEX library,
you would run the cellranger count pipeline:
$ cd /home/jdoe/runs $ cellranger count --id=GEX_VDJ_SampleGEX \ --transcriptome=/opt/refdata-cellranger-GRCh38-1.2.0 \ --fastqs=/home/jdoe/mkfastq_pipeline/outs/fastq_path \ --sample=SampleGEX \
This will generate a Loupe Cell Browser .cloupe file, a gene-barcode matrix, and other information.
For more details on running the cellranger count pipeline, consult the
Cell Ranger documentation.
Next, to analyze the data from the enriched T cell and B cell libraries, you will run the cellranger vdj pipeline:
$ cd /home/jdoe/runs $ cellranger vdj --id=GEX_VDJ_SampleT \ --reference=/opt/refdata-cellranger-vdj-GRCh38-alts-ensembl-2.0.0 \ --fastqs=/home/jdoe/mkfastq_pipeline/outs/fastq_path \ --sample=SampleT \
$ cd /home/jdoe/runs $ cellranger vdj --id=GEX_VDJ_SampleB \ --reference=/opt/refdata-cellranger-vdj-GRCh38-alts-ensembl-2.0.0 \ --fastqs=/home/jdoe/mkfastq_pipeline/outs/fastq_path \ --sample=SampleB \
These pipelines will generate a Loupe V(D)J Browser .vloupe file, in addition to a variety of files
to describe V(D)J clonotypes, consensus sequences, and contigs. For more details on running
the V(D)J analysis pipeline, consult the Cell Ranger V(D)J documentation.
The Loupe Cell Browser and Loupe V(D)J Browser software allow you to tie gene expression and V(D)J
information from these three pipeline runs back together. In Loupe Cell Browser 2.0, you can import
T Cell and B Cell V(D)J .vloupe files into the Loupe Cell Browser workspace, allowing you to explore
the immune repertoire within indvidual clusters, measure the gene expression of cells within
a clonotype of interest, and create new clusters based on V(D)J data:
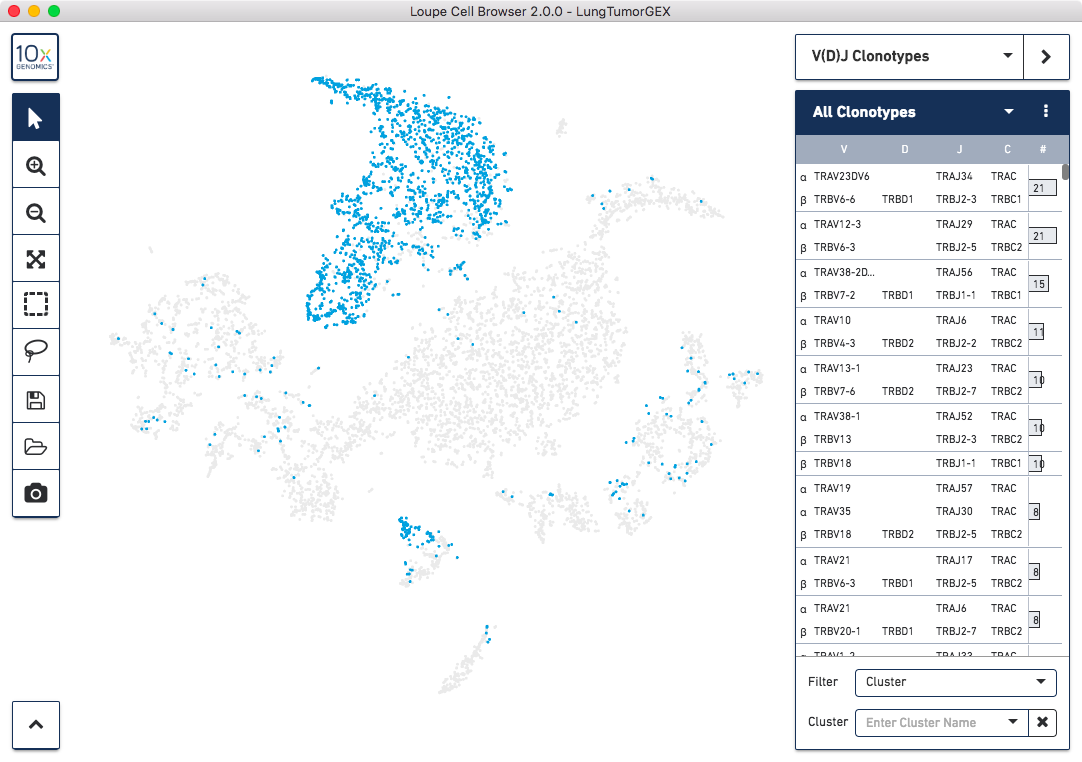
Click here for more information about how to use Loupe Cell Browser for gene expression and V(D)J analysis.
Likewise, you can import gene expression .cloupe files into Loupe V(D)J Browser, in order to
compare clonotypes between gene expression clusters, and explore V(D)J sequences within
clusters in greater detail:
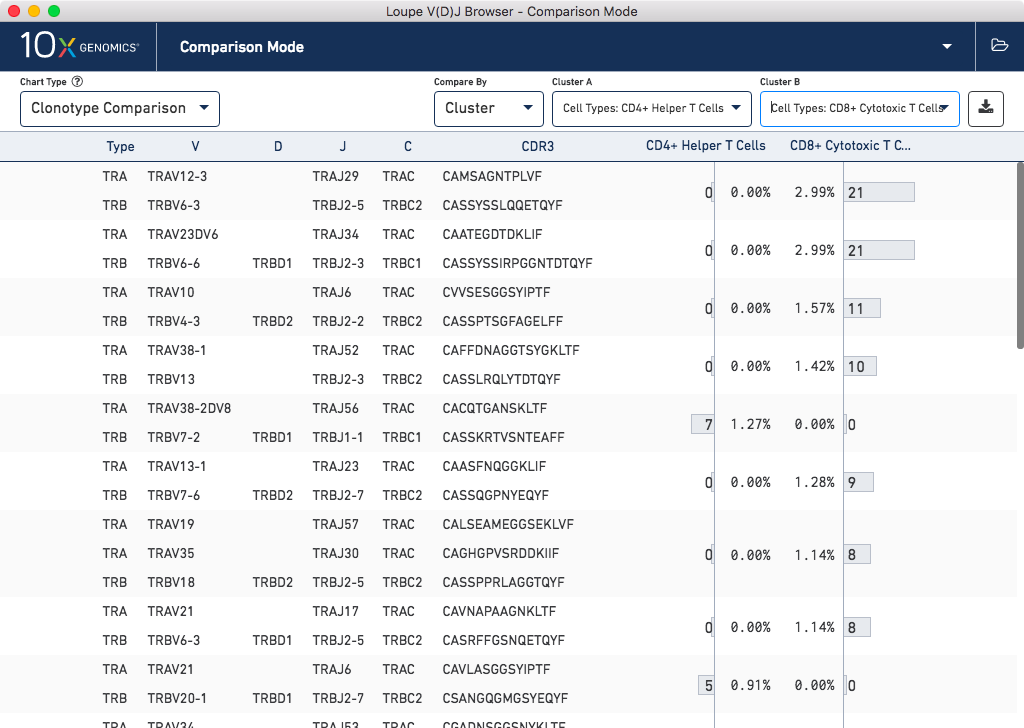
Click here for more information about how to use Loupe V(D)J Browser for more detailed analysis of V(D)J data.
To learn more about how to conduct multimodal analysis of V(D)J and gene expression data from the same sample, start the Loupe V(D)J + Gene Expression Tutorial. In the tutorial, you will be able to download example 5′ gene expression profiles and T cell clonotypes from a non-small cell lung carcinoma, and use the Loupe browsers to analyze the integrated data.